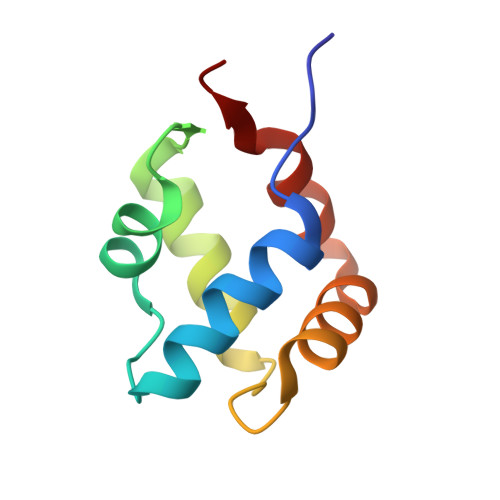
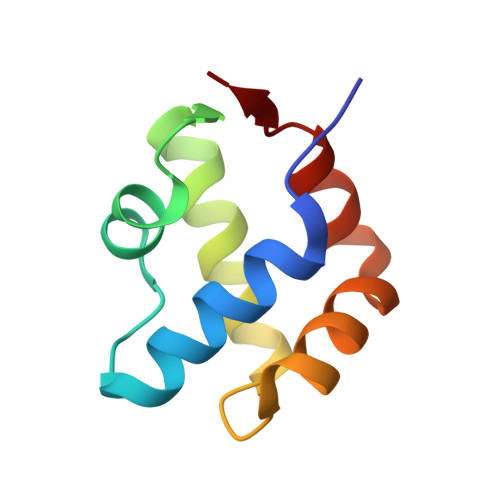
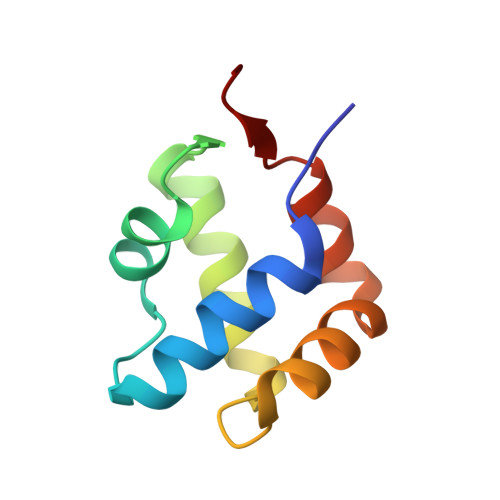
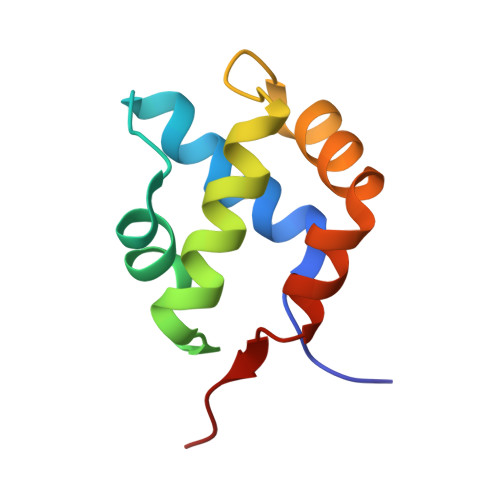
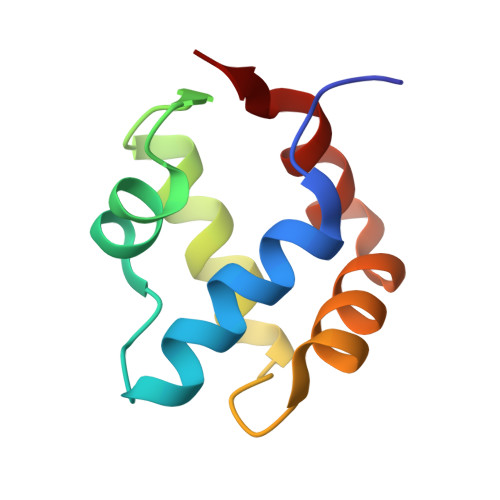
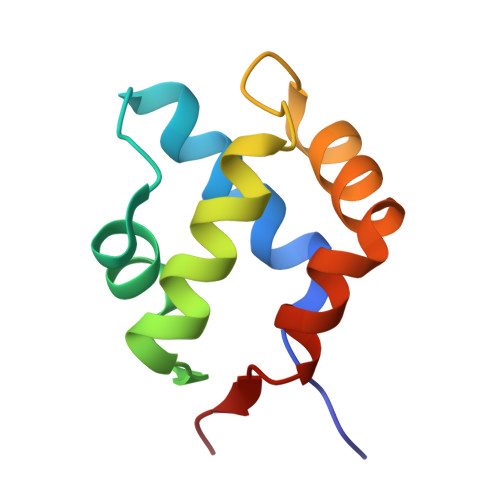
Characterization of an amyloid-based antiphage defence system in Escherichia coli.
Ibarlosa, L., Dheur, S., Sanchez, C., Deb, S., Granger-Farbos, A., Coustou, V., Berbon, M., Massoni-Laporte, A., Salin, B., Blancard, C., Saragaglia, C., Tene, X.M., Dufour, N., Rossier, O., Habenstein, B., Debarbieux, L., Kauffmann, B., Loquet, A., Saupe, S.J.(2025) Nat Microbiol 10: 2168-2178
- PubMed: 40813923
- DOI: https://doi.org/10.1038/s41564-025-02074-9
- Primary Citation of Related Structures:
9H9V - PubMed Abstract:
Amyloids are β-sheet-rich protein polymers that can cause disease but also serve functional purposes in cells. In animals and fungi, functional amyloids have a role in regulated cell death as molecular switches activating key cell death effectors. Here we describe an amyloid-based abortive infection antiphage defence system in Escherichia coli. This system leads to death of phage-infected cells and involves two proteins, Bab and Agp, which share a common amyloid motif and are encoded by adjacent genes. Following infection, Agp activates Bab through amyloid signalling. Activation of Bab then causes membrane alterations and cell death. We determined the structure of the cell death execution domain of Bab, which is distantly related to pore-forming domains found in fungi, animals and plants. We show that Bab and HET-S, a fungal amyloid-controlled regulated cell death execution protein from Podospora anserina, are functionally interchangeable. These findings show that amyloid-mediated immune signalling is conserved across kingdoms.
- IBGC, UMR 5095, CNRS-Université de Bordeaux, Bordeaux, France.
Organizational Affiliation: