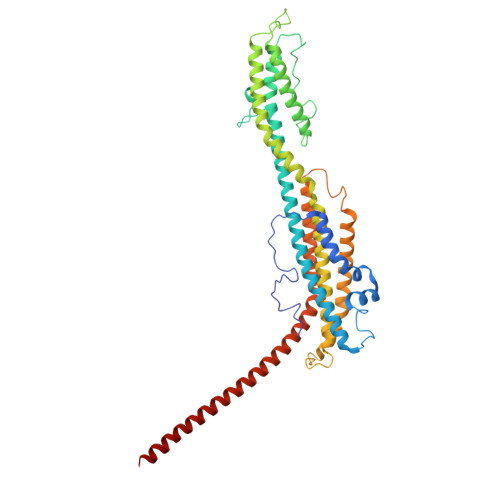
Cryo-EM structure and polar assembly of the PS2 S-layer of Corynebacterium glutamicum.
Sogues, A., Sleutel, M., Petit, J., Megrian, D., Bayan, N., Wehenkel, A.M., Remaut, H.(2025) Proc Natl Acad Sci U S A 122: e2426928122-e2426928122
- PubMed: 40729392 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2426928122
- Primary Citation Related Structures:
9GK2 - PubMed Abstract:
The polar-growing Corynebacteriales have a complex cell envelope architecture characterized by the presence of a specialized outer membrane composed of mycolic acids. In some Corynebacteriales , this mycomembrane is further supported by a proteinaceous surface layer or "S-layer," whose function, structure, and mode of assembly remain largely enigmatic. Here, we isolated ex vivo PS2 S-layers from the industrially important Corynebacterium glutamicum and determined its atomic structure by 3D cryo-EM reconstruction. PS2 monomers consist of a six-helix bundle "core," a three-helix bundle "arm," and a C-terminal transmembrane (TM) helix. The PS2 core oligomerizes into hexameric units anchored in the mycomembrane by a channel-like coiled-coil of the TM helices. The PS2 arms mediate trimeric lattice contacts, crystallizing the hexameric units into an intricate semipermeable lattice. Using pulse-chase live cell imaging, we show that the PS2 lattice is incorporated at the poles, coincident with the actinobacterial elongasome. Finally, phylogenetic analysis shows a paraphyletic distribution and dispersed chromosomal location of PS2 in Corynebacteriales as a result of multiple recombination events and losses. These findings expand our understanding of S-layer biology and enable applications of membrane-supported self-assembling bioengineered materials.
- Structural and Molecular Microbiology, Vlaams Instituut voor Biotechnology (VIB) Center for Structural Biology, Brussels 1050, Belgium.
Organizational Affiliation: