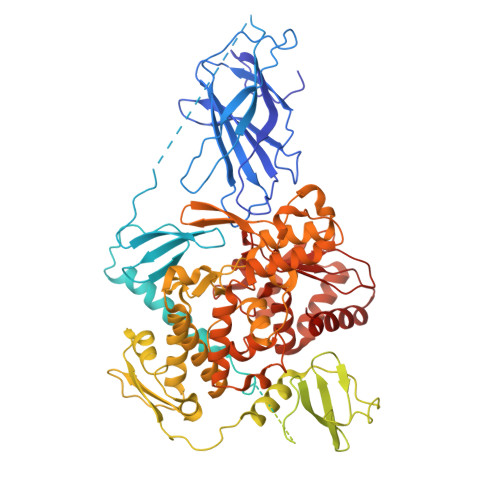
Structural basis of ubiquitin ligase Nedd4-2 autoinhibition and regulation by calcium and 14-3-3 proteins.
Janosev, M., Kosek, D., Tekel, A., Joshi, R., Honzejkova, K., Pohl, P., Obsil, T., Obsilova, V.(2025) Nat Commun 16: 4875-4875
- PubMed: 40419858 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-60207-4
- Primary Citation Related Structures:
9GIK, 9GIM - PubMed Abstract:
Nedd4-2 E3 ligase regulates Na + homeostasis by ubiquitinating various channels and membrane transporters, including the epithelial sodium channel ENaC. In turn, Nedd4-2 dysregulation leads to various conditions, including electrolytic imbalance, respiratory distress, hypertension, and kidney diseases. However, Nedd4-2 regulation remains mostly unclear. The present study aims at elucidating Nedd4-2 regulation by structurally characterizing Nedd4-2 and its complexes using several biophysical techniques. Our cryo-EM reconstruction shows that the C2 domain blocks the E2-binding surface of the HECT domain. This blockage, ubiquitin-binding exosite masking by the WW1 domain, catalytic C922 blockage and HECT domain stabilization provide the structural basis for Nedd4-2 autoinhibition. Furthermore, Ca 2+ -dependent C2 membrane binding disrupts C2/HECT interactions, but not Ca 2+ alone, whereas 14-3-3 protein binds to a flexible region of Nedd4-2 containing the WW2 and WW3 domains, thereby inhibiting its catalytic activity and membrane binding. Overall, our data provide key mechanistic insights into Nedd4-2 regulation toward fostering the development of strategies targeting Nedd4-2 function.
- Institute of Physiology of the Czech Academy of Sciences, Laboratory of Structural Biology of Signaling Proteins, Division BIOCEV, Prumyslova 595, 252 50, Vestec, Czech Republic.
Organizational Affiliation: