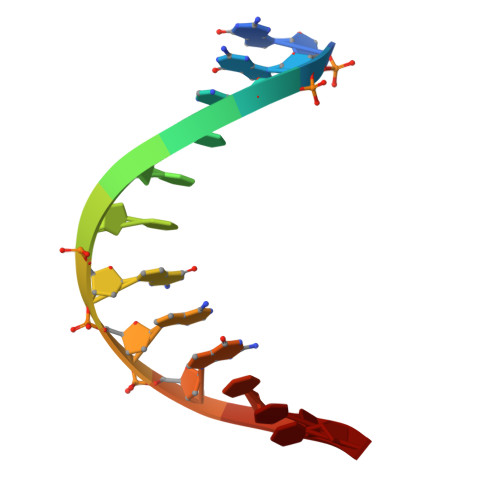
Unveiling the solution structure of a DNA duplex with continuous silver-modified Watson-Crick base pairs.
Javornik, U., Perez-Romero, A., Lopez-Chamorro, C., Smith, R.M., Dobado, J.A., Palacios, O., Bera, M.K., Nyman, M., Plavec, J., Galindo, M.A.(2024) Nat Commun 15: 7763-7763
- PubMed: 39237564 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-51876-8
- Primary Citation Related Structures:
9GDM - PubMed Abstract:
The challenge of transforming organized DNA structures into their metallized counterparts persists in the scientific field. In this context, utilizing DNA molecules modified with 7-deazapurine, provides a transformative solution. In this study, we present the solution structure of a DNA duplex that can be transformed into its metallized equivalent while retaining the natural base pairing arrangement through the creation of silver-modified Watson-Crick base pairs. Unlike previously documented X-ray structures, our research demonstrates the feasibility of preserving the intrinsic DNA self-assembly while incorporating Ag I into the double helix, illustrating that the binding of silver does not disrupt the canonical base-pairing organization. Moreover, in our case, the uninterrupted Ag I chain deviates from forming conventional straight linear chains; instead, it adheres to a helical arrangement dictated by the underlying DNA structure. This research challenges conventional assumptions and opens the door to precisely design structures based on the organization of highly stable Ag-DNA assemblies.
- Slovenian NMR Center, National Institute of Chemistry, SI-1000, Ljubljana, Slovenia.
Organizational Affiliation: