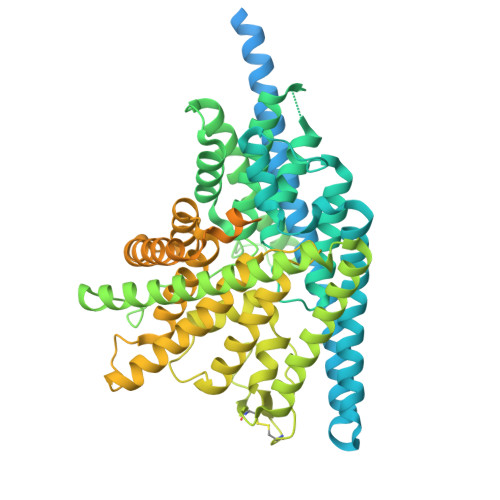
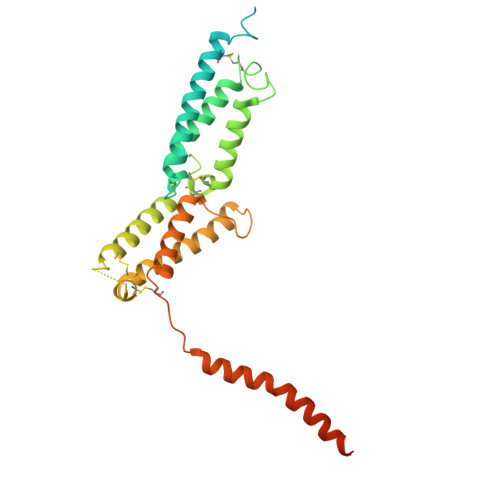
Mechanism of phosphoinositide regulation of lysosomal pH via inhibition of CLC-7.
Hilton, J.K., Lin, Y., Sefah, E., Deme, J.C., Parker, J.L., Langton, M.J., Grabe, M., Lea, S., Newstead, S., Mindell, J.A.(2025) bioRxiv
- PubMed: 41256531 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2025.10.01.679551
- Primary Citation Related Structures:
9G6C, 9G6D, 9G6E - PubMed Abstract:
Lysosomes process cellular waste and coordinate responses to metabolic challenge. Central to lysosomal homeostasis are phosphoinositide lipids, key signaling molecules which establish organelle identity, regulate membrane dynamics and are tightly linked to the pathophysiology and therapy of lysosomal storage disorders, neurodegeneration, and cancer. Phosphatidylinositol 3,5-bisphosphate (PI(3,5)P2) interacts with multiple lysosomal membrane proteins and plays a critical role in regulating lysosomal pH by directly inhibiting the chloride/proton antiporter ClC-7, though the molecular mechanism of this inhibition remains unclear. Here, using a combination of functional, structural, and computational analysis, we demonstrate that PI(3,5)P2 binding dramatically remodels the structure of ClC-7 by inducing close association between cytosolic and transmembrane domains. Disease-causing mutations show increased transport activity through loss of PI(3,5)P2 binding and subsequent inhibition. Conversely, ClC-7 activation is correlated with dissociation and increased disorder of the cytoplasmic domain along with novel transmembrane domain conformations, revealing a mechanistic link between specific lysosomal lipids, transporter regulation, and the enigmatic basis of the ClC-7 slow gate.
- Membrane Transport Biophysics Section, National Institute of Neurological Disorders and Stroke, NIH, Bethesda, MD 20892, USA.
Organizational Affiliation: