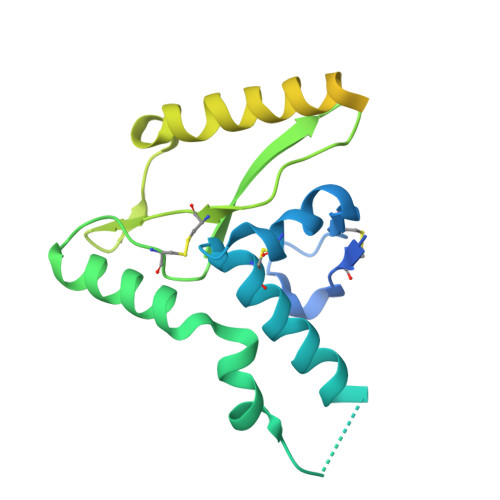
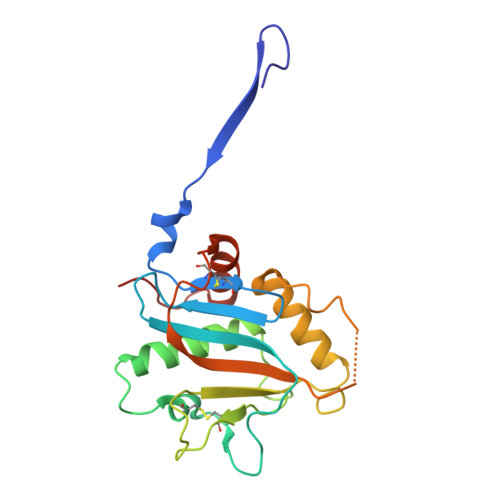
Metal-induced conformational changes in the Sabia virus spike complex.
Cohen-Dvashi, H., Katz, M., Diskin, R.(2025) Nat Microbiol 10: 2221-2230
- PubMed: 40751015 Search on PubMed
- DOI: https://doi.org/10.1038/s41564-025-02075-8
- Primary Citation Related Structures:
9FYA, 9FYE, 9FYG - PubMed Abstract:
Haemorrhagic fever viruses from the Arenaviridae are a source of concern owing to their potential to cause lethal outbreaks and the lack of effective therapeutics. While structures of spike proteins from 'Old World' arenaviruses are available, the differences and similarities to 'New World' arenaviruses, such as the Sabiá virus, remain unclear owing to the lack of New World spike structures. Here we present the structure of the isolated spike complex from the Sabiá virus, which mediates viral attachment and entry to the host cells, using single-particle cryo-electron microscopy. We find two distinct conformational states of the spike, representing its native closed state at 2.6 Å resolution and an open state at 2.9 Å resolution that it assumes during cell entry. In addition, we show that the opening of the spike and subsequent cell entry are dependent on acidic pH and an unidentified metal ion. Our study suggests potential differences in the cell entry mechanisms of clade B arenaviruses compared with others in the Arenaviridae family.
- Department of Chemical and Structural Biology, Weizmann Institute of Science, Rehovot, Israel.
Organizational Affiliation: