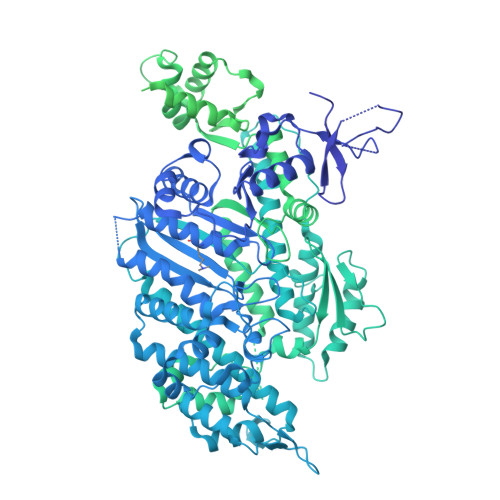
Development of clinically viable non-muscle myosin II small molecule inhibitors.
Radnai, L., Young, E.J., Kikuti, C., Toth, K., Zhou, M., Hafenbreidel, M., Stremel, R.F., Lin, L., Pasetto, P., Jin, X., Patel, A., Conlon, M., Briggs, S.B., Heidsieck, L., Sweeney, H.L., Sellers, J., Krieger-Burke, T., Martin, W.H., Sisco, J., Young, S., Pearson, P., Rumbaugh, G., Araldi, G.L., Duddy, S.K., Cameron, M.D., Surman, M., Houdusse, A., Griffin, P.R., Kamenecka, T.M., Miller, C.A.(2025) Cell 188: 4604-4621.e15
- PubMed: 40602401 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2025.06.006
- Primary Citation Related Structures:
9FU2 - PubMed Abstract:
Non-muscle myosin II (NMII), a molecular motor that regulates critical processes such as cytokinesis and neuronal plasticity, has substantial therapeutic potential. However, translating this potential to in vivo use has been hampered by a lack of selective tools. The most prototypical non-selective inhibitor inactivates both NMII and cardiac muscle myosin II (CMII), a key regulator of heart function. Using rational drug design, we developed a series of NMII inhibitors that markedly improve tolerability by selectively targeting NMII over CMII, including MT-228 and clinical candidate MT-110. MT-228 and MT-110 have excellent properties, including high brain penetration and efficacy in preclinical models of methamphetamine use disorder (MUD), which has no current FDA-approved therapies. The structure of MT-228 bound to myosin II provides insight into its selectivity for NMII over CMII. The broad therapeutic windows of these NMII inhibitors provide valuable tools for the scientific community and a promising clinical candidate for the treatment of MUD.
- Department of Molecular Medicine, The Scripps Research Institute and The Herbert Wertheim UF Scripps Institute for Biomedical Innovation & Technology, Jupiter, FL 33458, USA; Department of Neuroscience, The Scripps Research Institute and The Herbert Wertheim UF Scripps Institute for Biomedical Innovation & Technology, Jupiter, FL 33458, USA.
Organizational Affiliation: