Stapled Peptides as Inhibitors of mRNA Deadenylation.
Pal, S., Gordijenko, I., Schmeing, S., Biswas, S., Akbulut, Y., Gasper, R., 't Hart, P.(2025) Angew Chem Int Ed Engl 64: e202413911-e202413911
- PubMed: 39319385 Search on PubMed
- DOI: https://doi.org/10.1002/anie.202413911
- Primary Citation Related Structures:
9FL8 - PubMed Abstract:
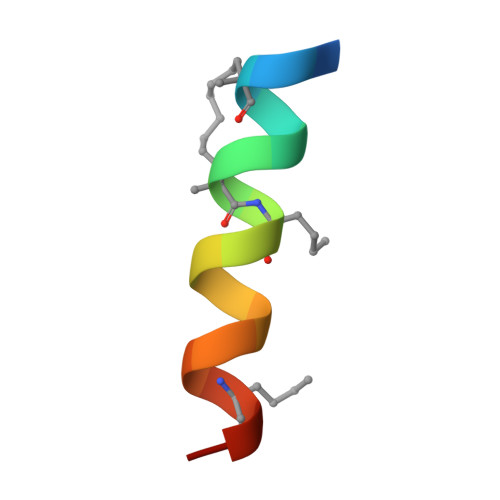
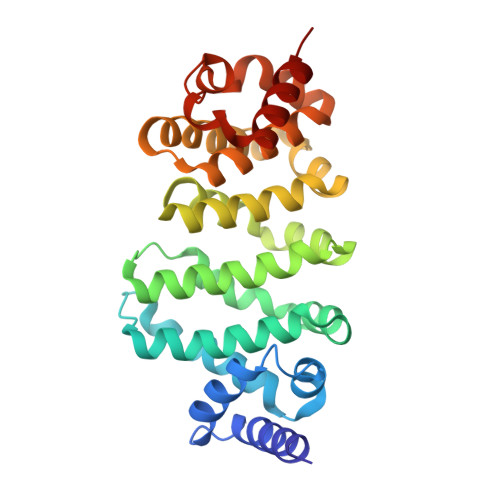
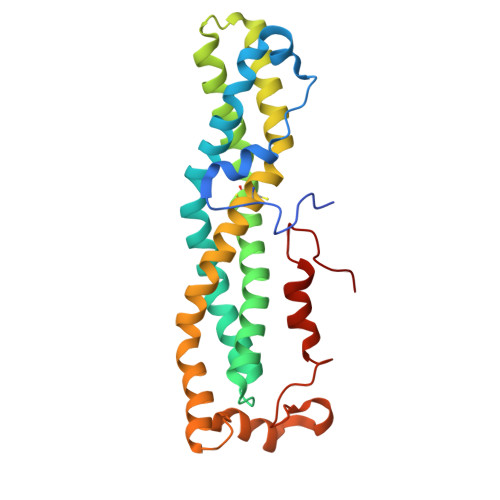
Therapeutic intervention targeting mRNA typically aims at reducing the levels of disease-causing sequences. Achieving the opposite effect of blocking the destruction of beneficial mRNA remains underexplored. The degradation of mRNA starts with the removal of poly(A) tails, reducing their stability and translational activity, which is mainly regulated by the CCR4-NOT complex. The subunit NOT9 binds various RNA binding proteins, that recruit mRNA in a sequence-specific manner to the CCR4-NOT complex to promote their deadenylation. These RNA binding proteins interact with NOT9 through a helical NOT9 binding motif, which we used as a starting point for development of the hydrocarbon stapled peptide NIP-2. The peptide (K D =60.4 nM) was able to inhibit RNA-binding (IC 50 =333 nM) as well as the deadenylation activity of the CCR4-NOT complex in vitro while being cell-permeable (cell-permeability EC 50 =2.44 μM). A co-crystal structure of NIP-2 bound to NOT9 allowed further optimization of the peptide through point mutation leading to NIP-2-H27A-N 3 (K D =122 nM) with high cell permeability (cell-permeability EC 50 =0.34 μM). The optimized peptide was able to inhibit deadenylation of target mRNAs when used in HeLa cells at a concentration of 100 μM, demonstrating the feasibility of increasing mRNA stability.
- Chemical Genomics Centre, Max Planck Institute of Molecular Physiology, Otto-Hahn-Strasse 11, 44227, Dortmund, Germany.
Organizational Affiliation: