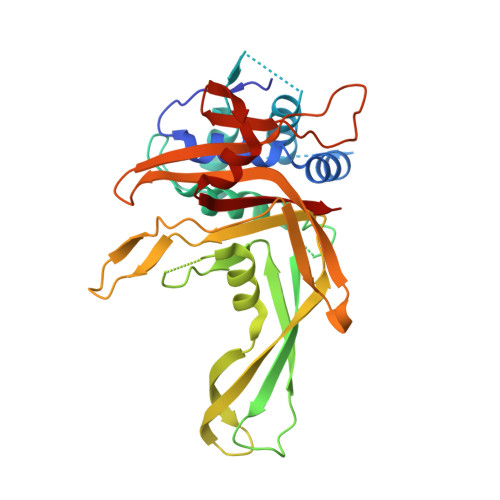
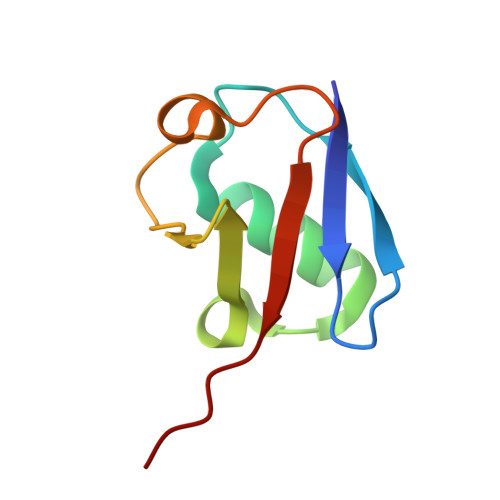
Chimeric deubiquitinase engineering reveals structural basis for specific inhibition of the mitophagy regulator USP30.
Kazi, N.H., Klink, N., Gallant, K., Kipka, G.M., Gersch, M.(2025) Nat Struct Mol Biol 32: 1776-1786
- PubMed: 40325251 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-025-01534-4
- Primary Citation Related Structures:
9F19, 9F6G - PubMed Abstract:
The mitochondrial deubiquitinase ubiquitin-specific protease (USP) 30 negatively regulates PINK1-parkin-driven mitophagy. Whether enhanced mitochondrial quality control through inhibition of USP30 can protect dopaminergic neurons is currently being explored in a clinical trial for Parkinson's disease. However, the molecular basis for specific inhibition of USP30 by small molecules has remained elusive. Here we report the crystal structure of human USP30 in complex with a specific inhibitor, enabled by chimeric protein engineering. Our study uncovers how the inhibitor extends into a cryptic pocket facilitated by a compound-induced conformation of the USP30 switching loop. Our work underscores the potential of exploring induced pockets and conformational dynamics to obtain deubiquitinase inhibitors and identifies residues facilitating specific inhibition of USP30. More broadly, we delineate a conceptual framework for specific USP deubiquitinase inhibition based on a common ligandability hotspot in the Leu73 ubiquitin binding site and on diverse compound extensions. Collectively, our work establishes a generalizable chimeric protein-engineering strategy to aid deubiquitinase crystallization and enables structure-based drug design with relevance to neurodegeneration.
- Chemical Genomics Center, Max Planck Institute of Molecular Physiology, Dortmund, Germany.
Organizational Affiliation: