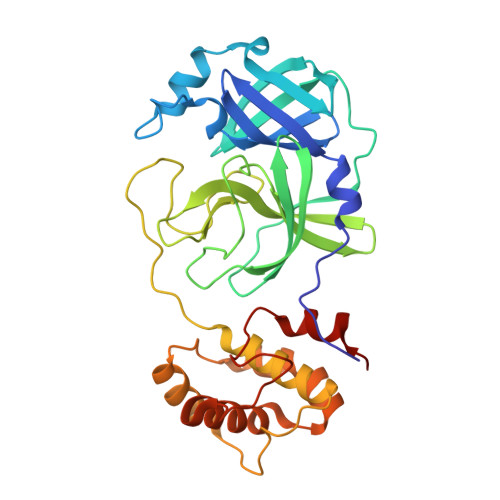
Allostery in homodimeric SARS-CoV-2 main protease.
Fornasier, E., Fabbian, S., Shehi, H., Enderle, J., Gatto, B., Volpin, D., Biondi, B., Bellanda, M., Giachin, G., Sosic, A., Battistutta, R.(2024) Commun Biol 7: 1435-1435
- PubMed: 39496839 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-024-07138-w
- Primary Citation Related Structures:
9EX8, 9EXU, 9EYA, 9EZ4, 9EZ6 - PubMed Abstract:
Many enzymes work as homodimers with two distant catalytic sites, but the reason for this choice is often not clear. For the main protease M pro of SARS-CoV-2, dimerization is essential for function and plays a regulatory role during the coronaviral replication process. Here, to analyze a possible allosteric mechanism, we use X-ray crystallography, native mass spectrometry, isothermal titration calorimetry, and activity assays to study the interaction of M pro with three peptide substrates. Crystal structures show how the plasticity of M pro is exploited to face differences in the sequences of the natural substrates. Importantly, unlike in the free form, the M pro dimer in complex with these peptides is asymmetric and the structures of the substrates nsp5/6 and nsp14/15 bound to a single subunit show allosteric communications between active sites. We identified arginines 4 and 298 as key elements in the transition from symmetric to asymmetric dimers. Kinetic data allowed the identification of positive cooperativity based on the increase in the processing efficiency (kinetic allostery) and not on the better binding of the substrates (thermodynamic allostery). At the physiological level, this allosteric behavior may be justified by the need to regulate the processing of viral polyproteins in time and space.
- Department of Chemical Sciences, University of Padova, via F. Marzolo 1, 35131, Padova, Italy.
Organizational Affiliation: