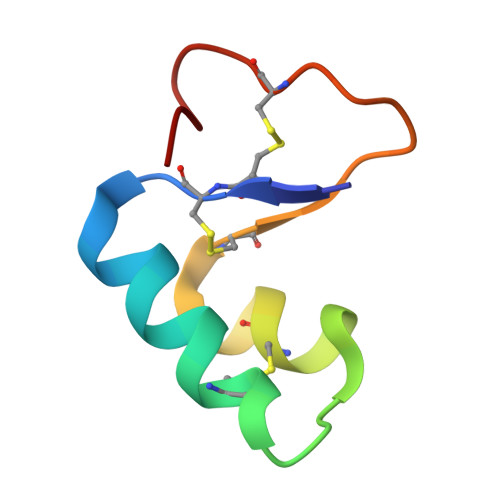
Solvent organization in the ultrahigh-resolution crystal structure of crambin at room temperature.
Chen, J.C.H., Gilski, M., Chang, C., Borek, D., Rosenbaum, G., Lavens, A., Otwinowski, Z., Kubicki, M., Dauter, Z., Jaskolski, M., Joachimiak, A.(2024) IUCrJ 11: 649-663
- PubMed: 39190507 Search on PubMed
- DOI: https://doi.org/10.1107/S2052252524007784
- Primary Citation Related Structures:
9EWK - PubMed Abstract:
Ultrahigh-resolution structures provide unprecedented details about protein dynamics, hydrogen bonding and solvent networks. The reported 0.70 Å, room-temperature crystal structure of crambin is the highest-resolution ambient-temperature structure of a protein achieved to date. Sufficient data were collected to enable unrestrained refinement of the protein and associated solvent networks using SHELXL. Dynamic solvent networks resulting from alternative side-chain conformations and shifts in water positions are revealed, demonstrating that polypeptide flexibility and formation of clathrate-type structures at hydrophobic surfaces are the key features endowing crambin crystals with extraordinary diffraction power.
- Structural Biology Center, X-ray Science Division, Argonne National Laboratory, Lemont, IL 60439, USA.
Organizational Affiliation: