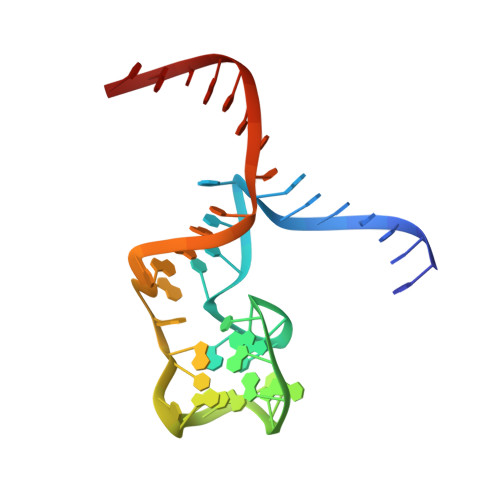
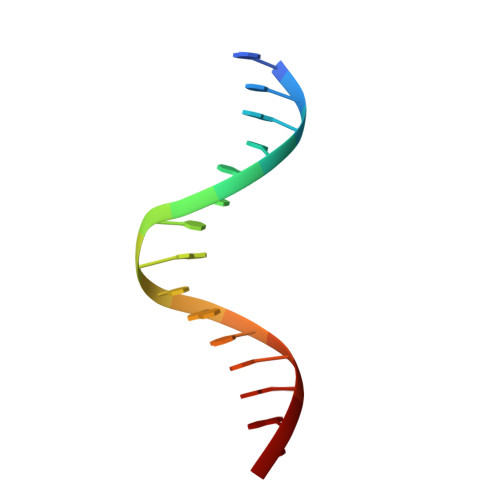
Designing Rigid DNA Origami Templates for Molecular Visualization Using Cryo-EM.
Khoshouei, A., Kempf, G., Mykhailiuk, V., Griessing, J.M., Honemann, M.N., Kater, L., Cavadini, S., Dietz, H.(2024) Nano Lett 24: 5031-5038
- PubMed: 38602296
- DOI: https://doi.org/10.1021/acs.nanolett.4c00915
- Primary Citation of Related Structures:
9EOQ - PubMed Abstract:
DNA origami, a method for constructing nanostructures from DNA, offers potential for diverse scientific and technological applications due to its ability to integrate various molecular functionalities in a programmable manner. In this study, we examined the impact of internal crossover distribution and the compositional uniformity of staple strands on the structure of multilayer DNA origami using cryogenic electron microscopy (cryo-EM) single-particle analysis. A refined DNA object was utilized as an alignment framework in a host-guest model, where we successfully resolved an 8 kDa thrombin binding aptamer (TBA) linked to the host object. Our results broaden the spectrum of DNA in structural applications.
- Laboratory for Biomolecular Nanotechnology, Department of Biosciences, School of Natural Sciences, Technical University of Munich, Am Coulombwall 4a, 85748 Garching, Germany.
Organizational Affiliation: