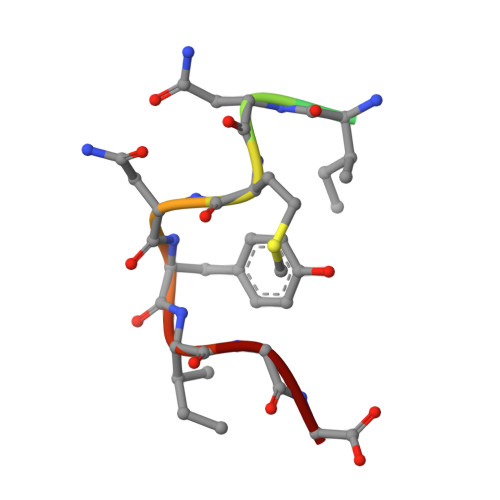
Mechanism of controlled radical initiation in radical SAM GTP 3',8-cyclase.
Pang, H., Li, D., Wu, Q., Zhang, P., Yang, W., Silakov, A., Zhou, P., Yokoyama, K.(2025) Proc Natl Acad Sci U S A 122: e2502098122-e2502098122
- PubMed: 41183211
- DOI: https://doi.org/10.1073/pnas.2502098122
- Primary Citation Related Structures:
9EFO - PubMed Abstract:
Metalloenzymes couple substrate binding and formation of oxidative intermediates to minimize unwanted side reactions. However, the molecular details of such coupling frequently remain ambiguous. Radical S -adenosyl-L-methionine (SAM) enzymes constitute one of the largest groups of metalloenzymes and catalyze various radical-mediated reactions. While radical SAM enzymes significantly accelerate the conserved radical initiation reaction, the reductive cleavage of SAM into 5'-deoxyadenosyl radical (5'-dA•), the molecular mechanism of this rate acceleration is largely unexplored. Here, using MoaA, aradical SAM enzyme in the molybdenum cofactor (Moco) biosynthesis, as a model, we reveal the mechanism of substrate-triggered radical initiation. We first elucidated the intact active site structure of MoaA using solution NMR characterization of the C-terminal tail, which was disordered in the reported crystal structures, and its computational docking into the MoaA structure. Together with the comprehensive functional validation, we show that MoaA uses its conformationally flexible C-terminal tail with two conserved Gly residues (GG motif) at the C-terminus as a sensor to detect substrate guanosine 5'-triphosphate (GTP) binding and trigger reductive SAM cleavage. Importantly, mutations that disrupt this regulatory mechanism cause Moco deficiency disease in humans. Comparison of these observations with other radical SAM enzymes provides insight into the general mechanism of substrate-triggered radical initiation in radical SAM enzymes.
- Department of Biochemistry, Duke University School of Medicine, Durham, NC 27710.
Organizational Affiliation: