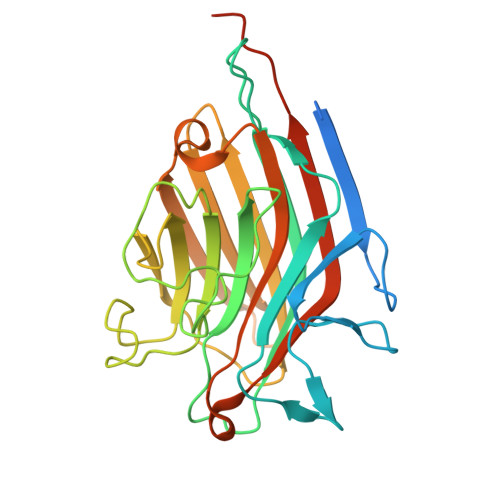
Maackia amurensis seed lectin structure and sequence comparison with other M. amurensis lectins.
Nayak, A.R., Holdcraft, C.J., Yin, A.C., Nicoletto, R.E., Zhao, C., Zheng, H., Temiakov, D., Goldberg, G.S.(2025) J Biological Chem 301: 108466-108466
- PubMed: 40158854 Search on PubMed
- DOI: https://doi.org/10.1016/j.jbc.2025.108466
- Primary Citation Related Structures:
9E6H - PubMed Abstract:
Maackia amurensis lectins, including MASL, MAA, and MAL2, are widely utilized in biochemical and medicinal research. However, the structural and functional differences between these lectins have not been defined. Here, we present a high-resolution cryo-EM structure of MASL revealing that its tetrameric assembly is directed by two intersubunit disulfide bridges. These bridges, formed by C272 residues, are central to the dimer-of-dimers assembly of a MASL tetramer. This cryo-EM structure also identifies residues involved in stabilizing the dimer interface, multiple glycosylation sites, and calcium and manganese atoms in the sugar-binding pockets of MASL. Notably, our analysis reveals that Y250 in the carbohydrate-binding site of MASL adopts a flipped conformation, likely acting as a gatekeeper that obstructs access to non-cognate substrates, a feature that may contribute to MASL's substrate specificity. Sequence analysis suggests that MAA is a truncated version of MASL, while MAL2 represents a homologous isoform. Unlike MASL, neither MAL2 nor MAA contains a cysteine residue required for disulfide bridge formation. Accordingly, analysis of these proteins using reducing and nonreducing SDS-PAGE confirms that the C272 residue in MASL drives intermolecular disulfide bridge formation. These findings provide critical insights into the unique structural features of MASL that distinguish it from other Maackia amurensis lectins, offering a foundation for further exploration of its biological and therapeutic potential.
- Biochemistry & Molecular Biology Department, Thomas Jefferson University, 1020 Locust Street, Philadelphia, PA 19107, USA.
Organizational Affiliation: