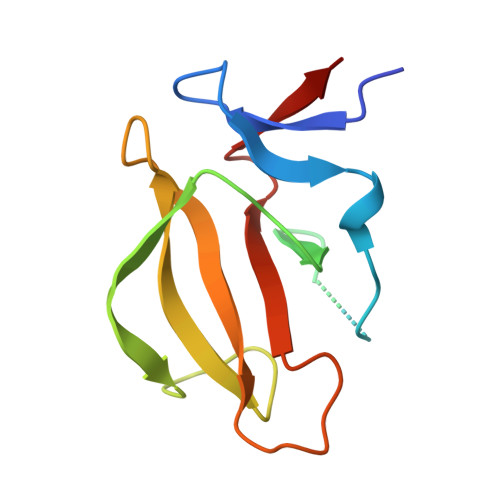
Discovery of KT-474─a Potent, Selective, and Orally Bioavailable IRAK4 Degrader for the Treatment of Autoimmune Diseases.
Zheng, X., Ji, N., Campbell, V., Slavin, A., Zhu, X., Chen, D., Rong, H., Enerson, B., Mayo, M., Sharma, K., Browne, C.M., Klaus, C.R., Li, H., Massa, G., McDonald, A.A., Shi, Y., Sintchak, M., Skouras, S., Walther, D.M., Yuan, K., Zhang, Y., Kelleher, J., Liu, G., Luo, X., Mainolfi, N., Weiss, M.M.(2024) J Med Chem 67: 18022-18037
- PubMed: 39151120 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.4c01305
- Primary Citation Related Structures:
9CUO - PubMed Abstract:
Interleukin-1 receptor associated kinase 4 (IRAK4) is an essential mediator of the IL-1R and TLR signaling pathways, both of which have been implicated in multiple autoimmune conditions. Hence, blocking the activity of IRAK4 represents an attractive approach for the treatment of autoimmune diseases. The activity of this serine/threonine kinase is dependent on its kinase and scaffolding activities; thus, degradation represents a potentially superior approach to inhibition. Herein, we detail the exploration of structure-activity relationships that ultimately led to the identification of KT-474, a potent, selective, and orally bioavailable heterobifunctional IRAK4 degrader. This represents the first heterobifunctional degrader evaluated in a nononcology indication and dosed to healthy human volunteers. This molecule successfully completed phase I studies in healthy adult volunteers and patients with atopic dermatitis or hidradenitis suppurativa. Phase II clinical trials in both of these indications have been initiated.
- Kymera Therapeutics, 500 N. Beacon Street, Fourth Floor, Watertown, Massachusetts 02472, United States.
Organizational Affiliation: