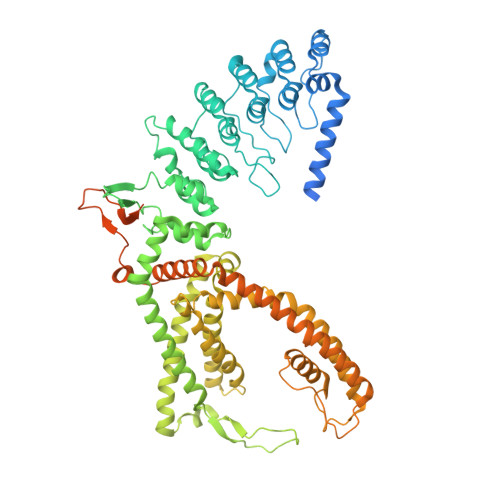
Structure-function analyses of human TRPV6 ancestral and derived haplotypes.
Neuberger, A., Shalygin, A., Trofimov, Y.A., Veretenenko, I.I., Nadezhdin, K.D., Krylov, N.A., Gudermann, T., Efremov, R.G., Chubanov, V., Sobolevsky, A.I.(2025) Structure 33: 91-103.e5
- PubMed: 39500315 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2024.10.018
- Primary Citation Related Structures:
9CUH, 9CUI, 9CUJ, 9CUK - PubMed Abstract:
TRPV6 is a Ca 2+ selective channel that mediates calcium uptake in the gut and contributes to the development and progression of human cancers. TRPV6 is represented by the ancestral and derived haplotypes that differ by three non-synonymous polymorphisms, located in the N-terminal ankyrin repeat domain (C157R), S1-S2 extracellular loop (M378V), and C-terminus (M681T). The ancestral and derived haplotypes were proposed to serve as genomic factors causing a different outcome for cancer patients of African ancestry. We solved cryoelectron microscopy (cryo-EM) structures of ancestral and derived TRPV6 in the open and calmodulin (CaM)-bound inactivated states. Neither state shows substantial structural differences caused by the non-synonymous polymorphisms. Functional properties assessed by electrophysiological recordings and Ca 2+ uptake measurements, and water and ion permeation evaluated by molecular modeling also appear similar between the haplotypes. Therefore, ancestral and derived TRPV6 have similar structure and function, implying that other factors are responsible for the differences in susceptibility to cancer.
- Department of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032, USA.
Organizational Affiliation: