Discovery of Daraxonrasib (RMC-6236), a Potent and Orally Bioavailable RAS(ON) Multi-selective, Noncovalent Tri-complex Inhibitor for the Treatment of Patients with Multiple RAS-Addicted Cancers.
Cregg, J., Edwards, A.V., Chang, S., Lee, B.J., Knox, J.E., Tomlinson, A.C.A., Marquez, A., Liu, Y., Freilich, R., Aay, N., Wang, Y., Jiang, L., Jiang, J., Wang, Z., Flagella, M., Wildes, D., Smith, J.A.M., Singh, M., Wang, Z., Gill, A.L., Koltun, E.S.(2025) J Med Chem 68: 6064-6083
- PubMed: 40056080 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.4c02314
- Primary Citation Related Structures:
9BG0, 9BG1, 9BG2, 9BG3, 9BG4, 9BG5, 9BG6, 9BG7, 9BG8, 9BG9, 9BGA, 9BGB, 9BGC, 9BGD - PubMed Abstract:
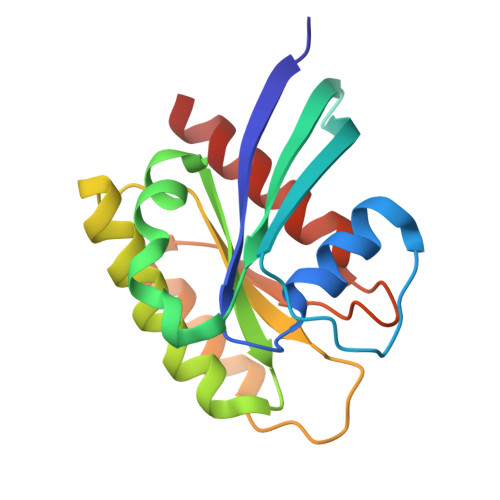
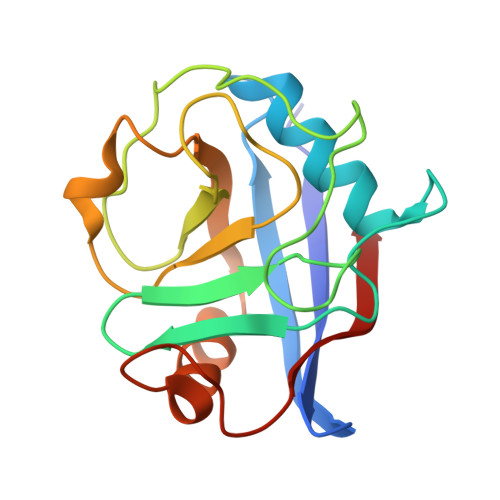
Oncogenic RAS mutations are among the most common in human cancers. To target the active, GTP-bound state of RAS(ON) directly, we employed an innovative tri-complex inhibitor (TCI) modality. Formation of a complex with an intracellular chaperone protein CypA, an inhibitor, and a target protein RAS blocks effector binding, inhibiting downstream RAS signaling and tumor cell proliferation. Herein, we describe the structure-guided SAR journey that led to the discovery of daraxonrasib (RMC-6236), a noncovalent, potent tri-complex inhibitor of multiple RAS mutant and wild-type (WT) variants. This orally bioavailable bRo5 macrocyclic molecule occupies a unique composite binding pocket comprising CypA and SWI/SWII regions of RAS(ON). To achieve broad-spectrum RAS isoform activity, we deployed an SAR campaign that focused on interactions with residues conserved between mutants and WT RAS isoforms. Concurrent optimization of potency and drug-like properties led to the discovery of daraxonrasib (RMC-6236), currently in clinical evaluation in RAS mutant advanced solid tumors (NCT05379985; NCT06040541; NCT06162221; NCT06445062; NCT06128551).
- Revolution Medicines, Inc., Redwood City, California 94063, United States.
Organizational Affiliation: