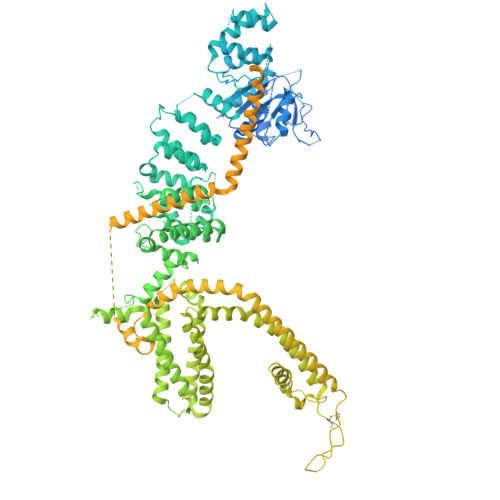
Molecular basis of neurosteroid and anticonvulsant regulation of TRPM3.
Yin, Y., Park, C.G., Feng, S., Guan, Z., Lee, H.J., Zhang, F., Sharma, K., Borgnia, M.J., Im, W., Lee, S.Y.(2025) Nat Struct Mol Biol 32: 828-840
- PubMed: 39809942 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-024-01463-8
- Primary Citation Related Structures:
9B28, 9B29, 9B2A - PubMed Abstract:
Transient receptor potential channel subfamily M member 3 (TRPM3) is a Ca 2+ -permeable cation channel activated by the neurosteroid pregnenolone sulfate (PregS) or heat, serving as a nociceptor in the peripheral sensory system. Recent discoveries of autosomal dominant neurodevelopmental disorders caused by gain-of-function mutations in TRPM3 highlight its role in the central nervous system. Notably, the TRPM3 inhibitor primidone, an anticonvulsant, has proven effective in treating patients with TRPM3-linked neurological disorders and in mouse models of thermal nociception. However, our understanding of neurosteroids, inhibitors and disease mutations on TRPM3 is limited. Here we present cryogenic electron microscopy structures of the mouse TRPM3 in complex with cholesteryl hemisuccinate, primidone and PregS with the synthetic agonist CIM 0216. Our studies identify the binding sites for the neurosteroid, synthetic agonist and inhibitor and offer insights into their effects and disease mutations on TRPM3 gating, aiding future drug development.
- Department of Biochemistry, Duke University School of Medicine, Durham, NC, USA.
Organizational Affiliation: