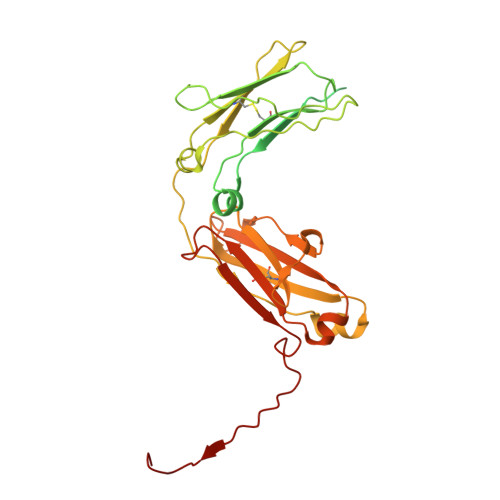
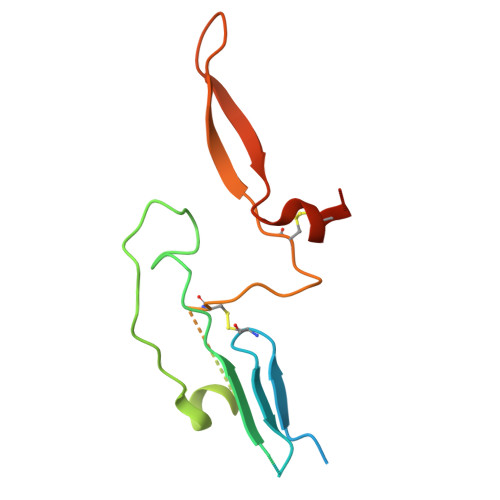
Adaptive multi-epitope targeting and avidity-enhanced nanobody platform for ultrapotent, durable antiviral therapy.
Xiang, Y., Xu, J., McGovern, B.L., Ranzenigo, A., Huang, W., Sang, Z., Shen, J., Diaz-Tapia, R., Pham, N.D., Teunissen, A.J.P., Rodriguez, M.L., Benjamin, J., Taylor, D.J., van Leent, M.M.T., White, K.M., Garcia-Sastre, A., Zhang, P., Shi, Y.(2024) Cell 187: 6966-6980.e23
- PubMed: 39447570 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2024.09.043
- Primary Citation Related Structures:
9ARV - PubMed Abstract:
Pathogens constantly evolve and can develop mutations that evade host immunity and treatment. Addressing these escape mechanisms requires targeting evolutionarily conserved vulnerabilities, as mutations in these regions often impose fitness costs. We introduce adaptive multi-epitope targeting with enhanced avidity (AMETA), a modular and multivalent nanobody platform that conjugates potent bispecific nanobodies to a human immunoglobulin M (IgM) scaffold. AMETA can display 20+ nanobodies, enabling superior avidity binding to multiple conserved and neutralizing epitopes. By leveraging multi-epitope SARS-CoV-2 nanobodies and structure-guided design, AMETA constructs exponentially enhance antiviral potency, surpassing monomeric nanobodies by over a million-fold. These constructs demonstrate ultrapotent, broad, and durable efficacy against pathogenic sarbecoviruses, including Omicron sublineages, with robust preclinical results. Structural analysis through cryoelectron microscopy and modeling has uncovered multiple antiviral mechanisms within a single construct. At picomolar to nanomolar concentrations, AMETA efficiently induces inter-spike and inter-virus cross-linking, promoting spike post-fusion and striking viral disarmament. AMETA's modularity enables rapid, cost-effective production and adaptation to evolving pathogens.
- Center of Protein Engineering and Therapeutics, Department of Pharmacological Sciences, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA.
Organizational Affiliation: