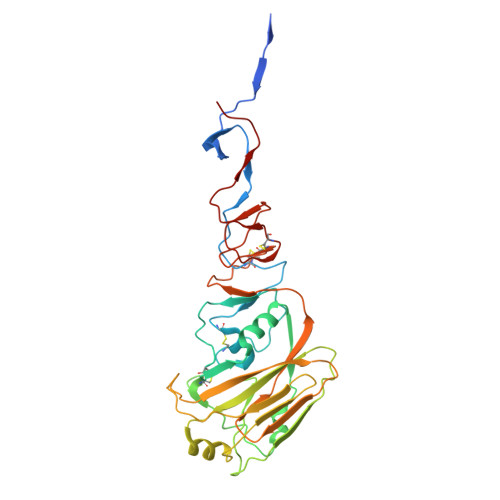
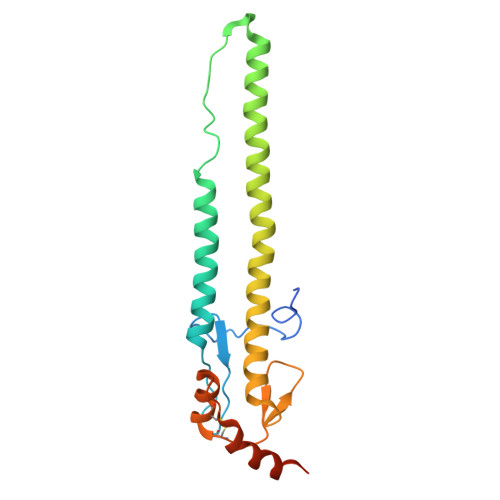
Multivalent interactions between fully glycosylated influenza virus hemagglutinins mediated by glycans at distinct N-glycosylation sites.
Li, R., Gao, J., Wang, L., Gui, M., Xiang, Y.(2024) Npj Viruses 2: 48-48
- PubMed: 40295773 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44298-024-00059-9
- Primary Citation Related Structures:
8ZDV, 8ZDW - PubMed Abstract:
The hemagglutinin (HA) glycoprotein of influenza virus binds host cell receptors and mediates viral entry. Here we present cryo-EM structures of fully glycosylated HAs from H5N1 and H5N8 influenza viruses. We find that the H5N1 HA can form filaments that comprise two head-to-head HA trimers. Multivalent interactions between the two HA trimers are mediated by glycans attached to N158. The distal Sia1-Gal2-NAG3 sugar moiety of N158 interacts with the receptor binding site on the opposing HA trimer. Additional interactions are observed between NAG3 and residues K222 and K193. The H5N8 HA lacks the N158 glycosylation site and does not form the filamentous structure. However, the H5N8 HA exhibits an auto-inhibition conformation, where the receptor binding site is occupied by the glycan chain attached to residue N169 from a neighboring protomer. These structures represent native HA-glycan interactions, which may closely mimic the receptor-HA interactions on the cell surface.
- Beijing Frontier Research Center for Biological Structure, Center for Infectious Disease Research, School of Basic Medical Sciences, Tsinghua University, 100084, Beijing, China.
Organizational Affiliation: