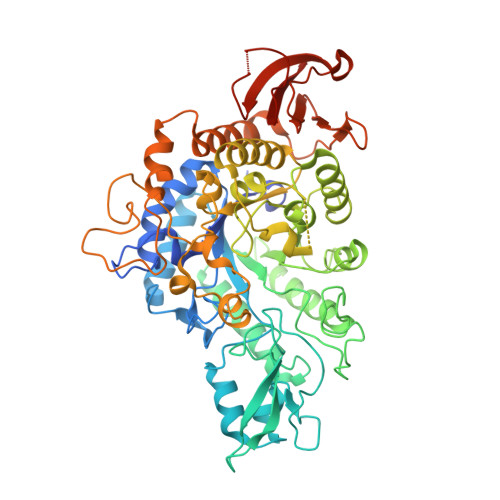
Molecular mechanism for the substrate specificity of Arthrobacter globiformis M6 alpha-glucosidase CmmB, belonging to glycoside hydrolase family 13 subfamily 30
Saburi, W., Tagami, T., Usui, T., Yu, J., Ose, T., Yao, M., Mori, H.(2024) Food Biosci 61: 104516