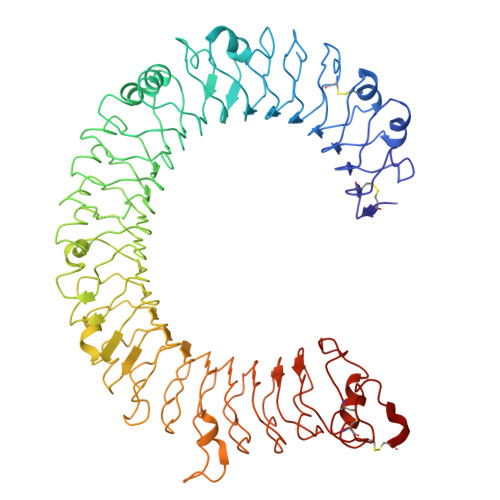
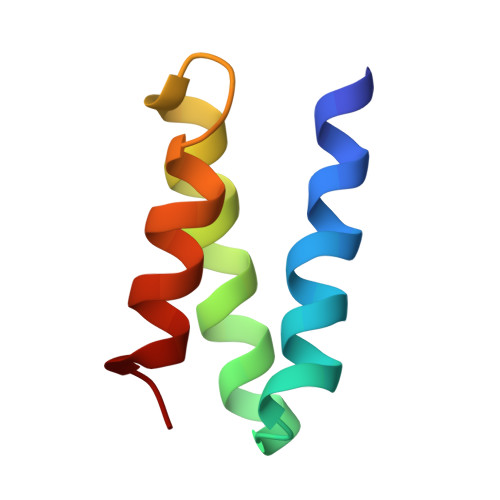
De novo design of protein minibinder agonists of TLR3.
Adams, C.S., Kim, H., Burtner, A.E., Lee, D.S., Dobbins, C., Criswell, C., Coventry, B., Tran-Pearson, A., Kim, H.M., King, N.P.(2025) Nat Commun 16: 1234-1234
- PubMed: 39890776 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-56369-w
- Primary Citation Related Structures:
8YHT, 8YHU - PubMed Abstract:
Toll-like Receptor 3 (TLR3) is a pattern recognition receptor that initiates antiviral immune responses upon binding double-stranded RNA (dsRNA). Several nucleic acid-based TLR3 agonists have been explored clinically as vaccine adjuvants in cancer and infectious disease, but present substantial manufacturing and formulation challenges. Here, we use computational protein design to create novel miniproteins that bind to human TLR3 with nanomolar affinities. Cryo-EM structures of two minibinders in complex with TLR3 reveal that they bind the target as designed, although one partially unfolds due to steric competition with a nearby N-linked glycan. Multivalent forms of both minibinders induce NF-κB signaling in TLR3-expressing cell lines, demonstrating that they may have therapeutically relevant biological activity. Our work provides a foundation for the development of specific, stable, and easy-to-formulate protein-based agonists of TLRs and other pattern recognition receptors.
- Institute for Protein Design, University of Washington, Seattle, WA, 98195, USA.
Organizational Affiliation: