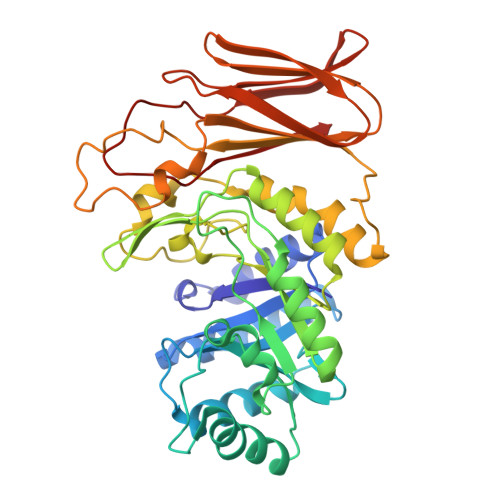
Crystal structure of GH71 alpha-1,3-glucanase Agn1p from Schizosaccharomyces pombe: an enzyme regulating cell division in fission yeast.
Horaguchi, Y., Saitoh, H., Konno, H., Makabe, K., Yano, S.(2025) Biochem Biophys Res Commun 766: 151907-151907
- PubMed: 40306164 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2025.151907
- Primary Citation Related Structures:
8YFH - PubMed Abstract:
Agn1p is a glycoside hydrolase family 71 α-1,3-glucanase from Schizosaccharomyces pombe. It is involved in cell division and releases nigero-pentaose from α-1,3-glucan as a primary hydrolysate. In this study, we used x-ray crystallography to determine the molecular structure of Agn1p, achieving a resolution of 1.80 Å for its free form and 2.10 Å for the substrate complex structure of an inactive mutant. We find that Agn1p comprises eight α-helices and sixteen β-strands, and these combined into a classical (α/β) 8 TIM-barrel core domain and a β-sandwich accessory domain. The TIM-barrel had a deep cavity in the center. Next, to determine which amino acid residues are involved in the catalytic reaction, we conducted substitution experiments on Asp-69, Asp-237, and Glu-240, three residues located in the cavity, preparing the corresponding substitution mutants D69N, D237A, D237N, E240A and E240Q. We found that the far-UV CD spectra of the five substitution mutants were similar to those of wild-type Agn1p, but all five mutants lost α-1,3-glucan hydrolyzing activity. We also obtained the cocrystal of the D237N mutant and nigero-heptaose, and its structure was determined. Specifically, we observed the electron density for the hexamer or pentamer sugar portion of nigero-heptaose. Moreover, the substrates were located in the vicinity of Asp-69, Asp-237, and Glu-240. Overall, these results suggest that Agn1p contains a stable substrate binding site for the hexamer or pentamer sugar structure of nigero-oligosaccharide.
- Graduate School of Sciences and Engineering, Yamagata UniversityJonan, Yonezawa, Yamagata, 992-8510, Japan.
Organizational Affiliation: