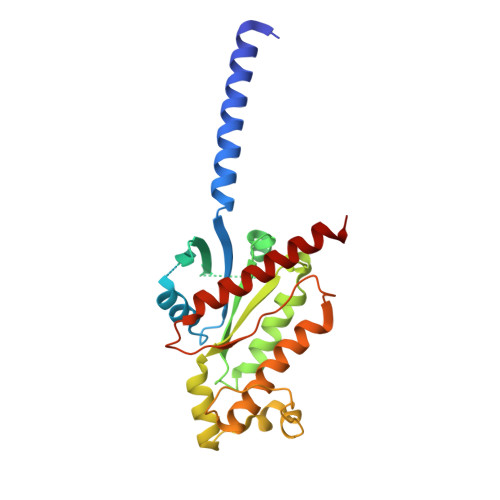
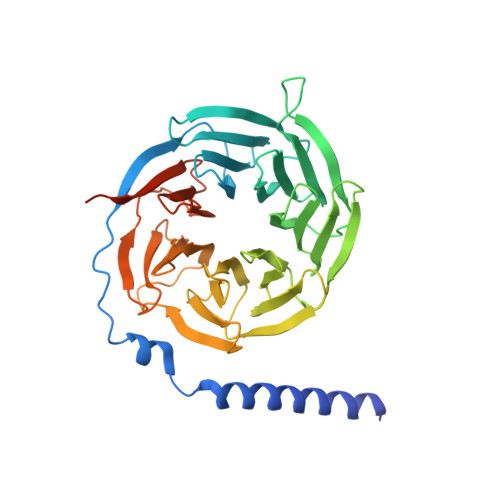
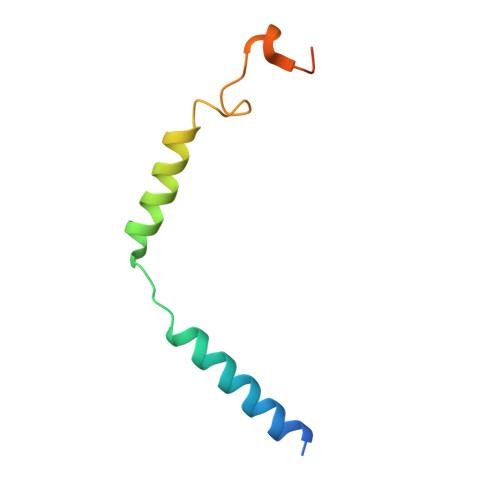
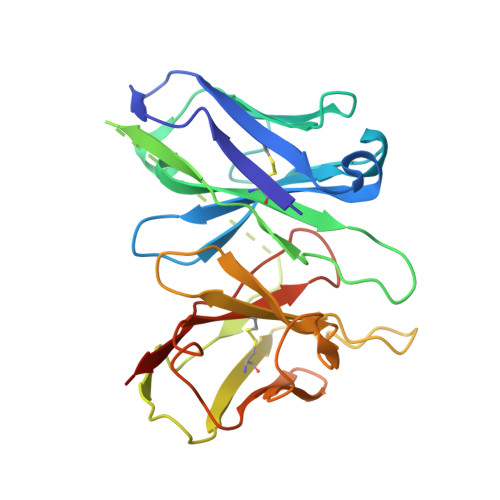
Structural basis for ligand recognition and activation of the prostanoid receptors.
Li, X., Zhang, X., Wen, X., Zhang, D., Qu, C., Miao, X., Zhang, W., Zhang, R., Liu, G., Xiao, P., Sun, J.P., Gong, W.(2024) Cell Rep 43: 113893-113893
- PubMed: 38446662 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2024.113893
- Primary Citation Related Structures:
8XJK, 8XJL, 8XJM, 8XJN, 8XJO - PubMed Abstract:
Prostaglandin F 2α (PGF 2α ) and thromboxane A 2 (TXA 2 ) are endogenous arachidonic acid metabolites, modulating diverse physiological processes including inflammation and cardiovascular homeostasis through activating PGF 2α receptor (FP) and TXA 2 receptor (TP). Ligands targeting FP and TP have demonstrated efficacy in treating conditions like glaucoma and cardiovascular diseases in humans, as well as reproductive-related diseases in animals. Here, we present five cryoelectron microscopy structures illustrating FP and TP in complex with G q and bound to PGF 2α (endogenous ligand), latanoprost acid (a clinical drug), and two other synthetic agonists. Combined with mutational and functional studies, these structures reveal not only structural features for the specific recognition of endogenous ligands and attainment of receptor selectivity of FP and TP but also the common mechanisms of receptor activation and G q protein coupling. The findings may enrich our knowledge of ligand recognition and signal transduction of the prostanoid receptor family and facilitate rational ligand design toward these two receptors.
- Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei, Anhui 230026, China.
Organizational Affiliation: