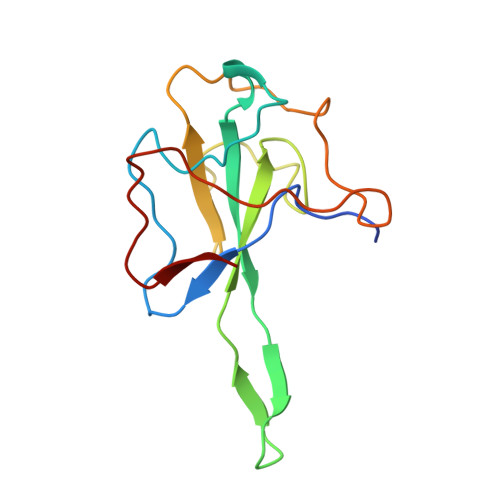
Unveiling potential inhibitors targeting the nucleocapsid protein of SARS-CoV-2: Structural insights into their binding sites.
Kumari, S., Mistry, H., Bihani, S.C., Mukherjee, S.P., Gupta, G.D.(2024) Int J Biol Macromol 273: 133167-133167
- PubMed: 38885868 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2024.133167
- Primary Citation Related Structures:
8X1H - PubMed Abstract:
The Nucleocapsid (N) protein of SARS-CoV-2 plays a crucial role in viral replication and pathogenesis, making it an attractive target for developing antiviral therapeutics. In this study, we used differential scanning fluorimetry to establish a high-throughput screening method for identifying high-affinity ligands of N-terminal domain of the N protein (N-NTD). We screened an FDA-approved drug library of 1813 compounds and identified 102 compounds interacting with N-NTD. The screened compounds were further investigated for their ability to inhibit the nucleic-acid binding activity of the N protein using electrophoretic mobility-shift assays. We have identified three inhibitors, Ceftazidime, Sennoside A, and Tannic acid, that disrupt the N protein's interaction with RNA probe. Ceftazidime and Sennoside A exhibited nano-molar range binding affinities with N protein, determined through surface plasmon resonance. The binding sites of Ceftazidime and Sennoside A were investigated using [ 1 H, 15 N]-heteronuclear single quantum coherence (HSQC) NMR spectroscopy. Ceftazidime and Sennoside A bind to the putative RNA binding site of the N protein, thus providing insights into the inhibitory mechanism of these compounds. These findings will contribute to the development of novel antiviral agents targeting the N protein of SARS-CoV-2.
- Protein Crystallography Section, Bhabha Atomic Research Centre, Mumbai, India; Homi Bhabha National Institute, Anushaktinagar, Mumbai, India.
Organizational Affiliation: