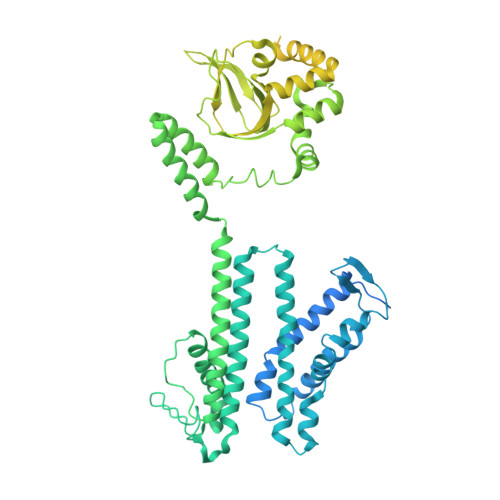
Structural changes in the conversion of an Arabidopsis outward-rectifying K + channel into an inward-rectifying channel.
Gao, X., Xu, X., Sun, T., Lu, Y., Jia, Y., Zhou, J., Fu, P., Zhang, Y., Yang, G.(2024) Plant Commun 5: 100844-100844
- PubMed: 38341617 Search on PubMed
- DOI: https://doi.org/10.1016/j.xplc.2024.100844
- Primary Citation Related Structures:
8WTZ, 8WUI - State Key Laboratory of Plant Environmental Resilience, Frontiers Science Center for Molecular Design Breeding, Department of Nutrition and Health, College of Biological Sciences, China Agricultural University, Beijing 100193, China.
Organizational Affiliation: