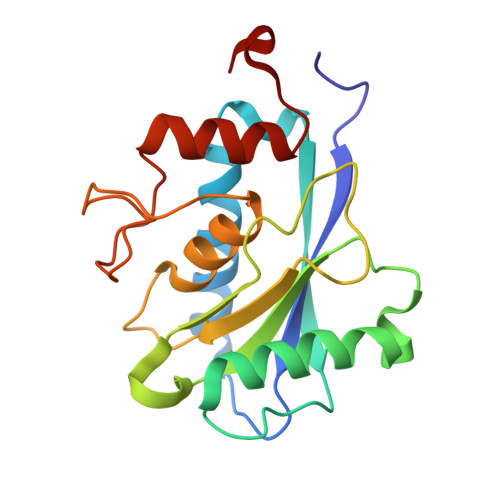
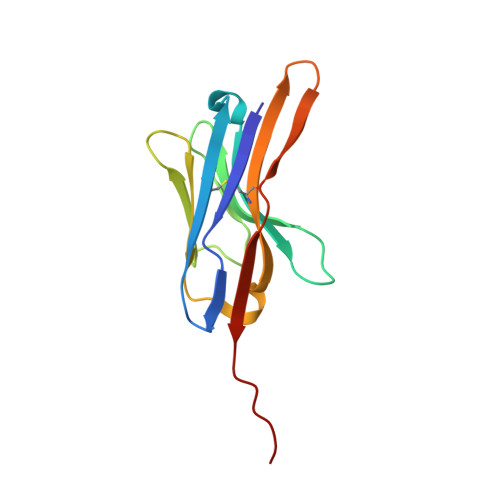
Mechanistic diversity of Bacteroides fragilis toxins and neutralization with single domain antibody.
Guo, Y., Ouyang, Z., He, W., Qin, Q., Zhang, J., Yu, T., Jiao, M., Hwang, P.M., Zheng, F., Muyldermans, S., Wen, Y.(2026) Cell Chem Biol 33: 102-116.e6
- PubMed: 41544614 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2025.12.009
- Primary Citation Related Structures:
8WEM, 8WEN, 8WEO - PubMed Abstract:
Enterotoxigenic Bacteroides fragilis (ETBF) promotes colonic inflammation by secreting metalloenzyme toxins (BFTs). Understanding BFT mechanisms and developing neutralization strategies is critical. Here, we have solved the structures of BFT-1 and BFT-2, revealing that residue 357 in the active site of the catalytic domain explains the diversity of function observed in BFT subtypes. We demonstrate that BFTs can directly cleave human epithelial-cadherin at extracellular domain 4, with BFT-2 possessing the highest activity. Using an alpaca antibody library, we identified a single-domain antibody, Nb2.43, targeting the BFTs. Nb2.43 can neutralize all three subtypes of BFT by directly binding the metalloenzyme catalytic zinc ion with its CDR3 antigen-binding loop. Furthermore, Nb2.43 blocks cleavage of E-cadherin by BFT and prevents the damage caused by ETBF in vitro and in a mouse colitis model. This work provides structural insights into BFT diversity and delivers a therapeutic nanobody against ETBF-mediated inflammation.
- Center for Microbiome Research of Med-X Institute, Department of Critical Care Medicine, Shaanxi Provincial Key Laboratory of Sepsis in Critical Care Medicine, The First Affiliated Hospital, Xi'an Jiaotong University, Xi'an 710061, China; The Key Laboratory of Environment and Genes Related to Disease of Ministry of Education, Health Science Center, Basic Medical Science, Xi'an Jiaotong University, Xi'an 710061, China; School of Basic Medical Sciences, Cheeloo College of Medicine, Shandong University, Jinan, China.
Organizational Affiliation: