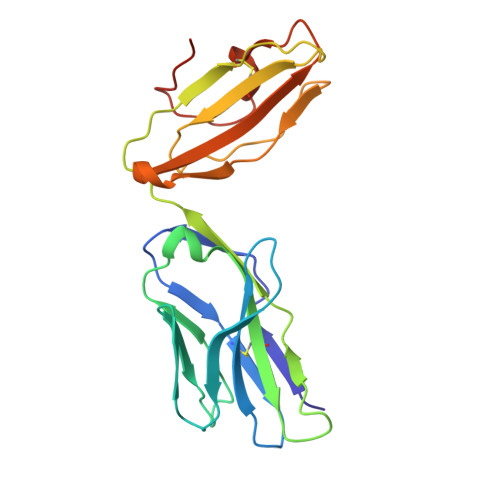
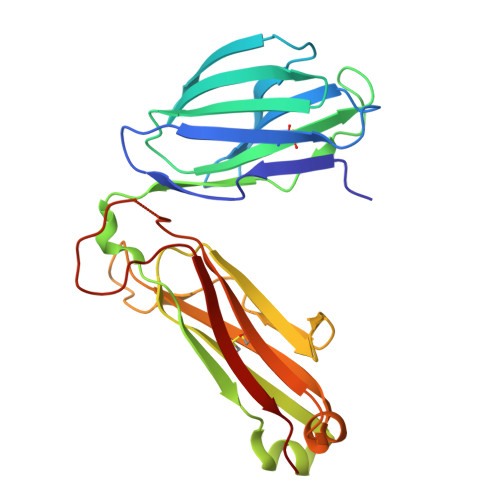
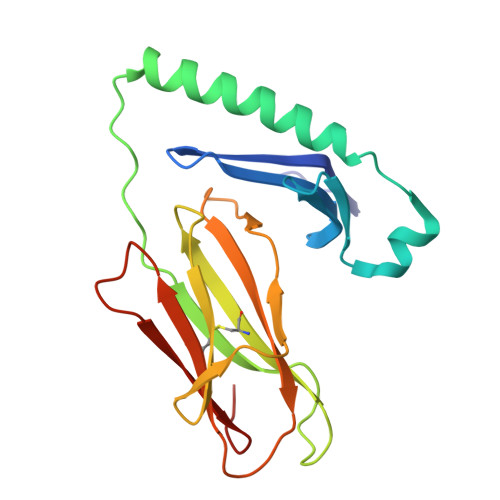
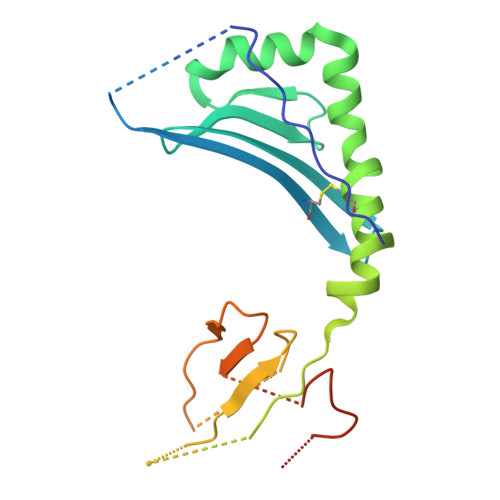
LCK-co-receptor association ensures T cell lineage fidelity and maximizes epitope-specific TCR diversity.
Zhang, J.B., Chaurasia, P., Nguyen, A., Huang, Z., Nguyen, T.T., Xu, H., Tran, M.T., Reid, H.H., Jones, C.M., Schattgen, S.A., Thiele, D., Thomas, P.G., Rientjes, J., Good-Jacobson, K.L., Ruscher, R., Littler, D.R., Rossjohn, J., Zareie, P., La Gruta, N.L.(2025) Sci Immunol 10: eadp5016-eadp5016
- PubMed: 39982976 Search on PubMed
- DOI: https://doi.org/10.1126/sciimmunol.adp5016
- Primary Citation Related Structures:
8VQ8, 9AUD - PubMed Abstract:
The interaction between the CD4/CD8 co-receptors and LCK (an Src family tyrosine kinase) is thought to augment T cell activation upon recognition of peptide-loaded major histocompatibility complexes (pMHCs). How this interaction influences antigen-specific T cell development is unclear however, as is its impact on naïve and immune antigen-specific T cell repertoires. In mice expressing mutated endogenous LCK unable to bind co-receptors (LCK FREE mice), we show that influenza A virus (IAV)-derived pMHC-specific CD8 and CD4 T cell responses had a significantly narrowed T cell receptor (TCR) repertoire, favoring high-affinity TCRs. This narrowing was established during T cell development and was exacerbated after viral infection. The dissociation of LCK from co-receptors also resulted in the redirection of CD4-fated T cells to the CD8 lineage, with expanded pMHCII-specific cytotoxic CD8 T cells observed after IAV infection. Thus, LCK-co-receptor association is critical for ensuring T cell lineage fidelity and maximizing antigen-specific T cell repertoire diversity.
- Immunity Program and Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, VIC, Australia.
Organizational Affiliation: