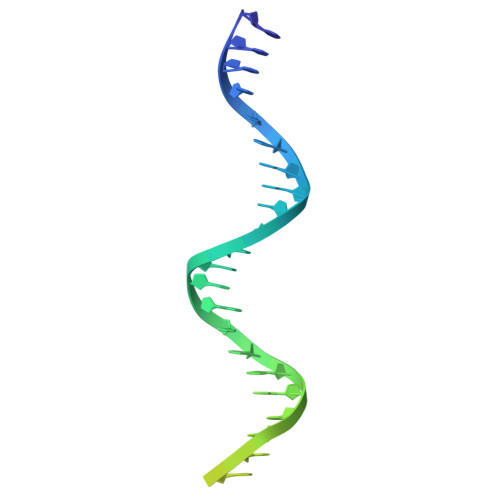
Bringing the ends together: cryo-EM structures of mycobacterial Ku in complex with DNA define its role in NHEJ synapsis.
Baral, J., Ang, C.S., McMillan, P.J., Shobhana, K., Saini, A., Hinde, E., Das, A.K., Rouiller, I.(2026) Nucleic Acids Res 54
- PubMed: 41521670 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaf1418
- Primary Citation Related Structures:
8V53, 8VF2, 8VF4, 8VF5 - PubMed Abstract:
Non-homologous end joining (NHEJ) is the sole pathway for repairing double-strand breaks in Mycobacterium tuberculosis during dormancy, relying on mycobacterial Ku (mKu) and ligase D, with mKu as the rate-limiting factor. Despite its essential role, the lack of structural information on prokaryotic Ku has hindered understanding of the molecular mechanisms underlying bacterial two-component NHEJ machinery. Here, we present the first cryo-electron microscopy (cryo-EM) structures of mKu in DNA-bound and higher-order supercomplex forms, revealing a Ku-mediated DNA synapsis mechanism unique to prokaryotes. Integrating cryo-EM with hydrogen-deuterium exchange mass spectrometry, we define key mKu-mKu dimerization, DNA-binding, and synapsis interactions essential for efficient NHEJ, bridging structure with function. Structure-guided in silico mutagenesis, coupled with electrophoretic mobility shift assays, identifies residues essential for DNA binding and synaptic assembly, which are crucial for NHEJ. Förster resonance energy transfer confirms DNA-dependent mKu oligomerization in solution, while live-cell imaging captures its spatiotemporal dynamics during double-stranded DNA break repair. These findings provide fundamental insights into the architecture and function of prokaryotic NHEJ, positioning mKu as a potential therapeutic target against tuberculosis and offering a framework for understanding DNA repair across bacterial species.
- Department of Bioscience and Biotechnology, Indian Institute of Technology Kharagpur, West Midnapore 721302, West Bengal, India.
Organizational Affiliation: