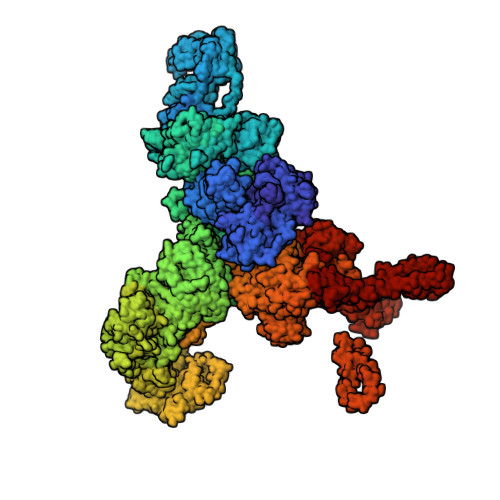
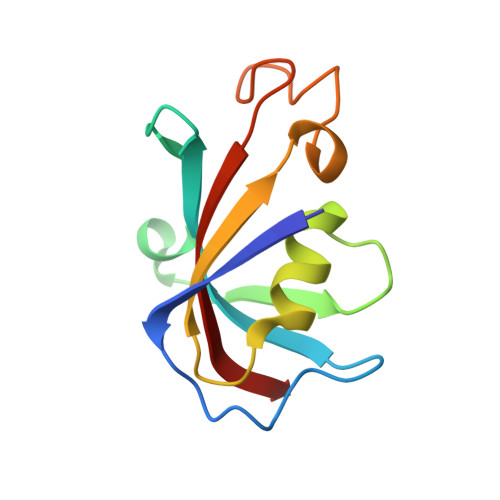
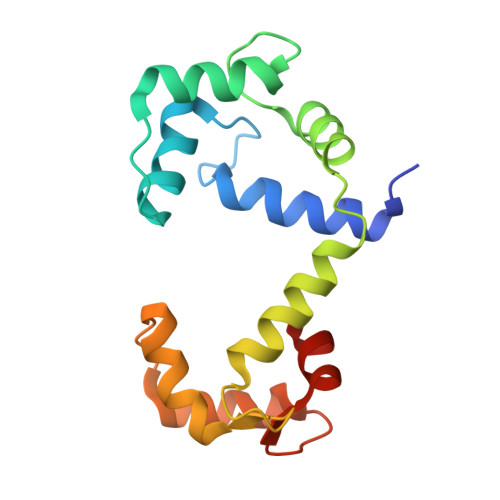
Structural basis for ryanodine receptor type 2 leak in heart failure and arrhythmogenic disorders.
Miotto, M.C., Reiken, S., Wronska, A., Yuan, Q., Dridi, H., Liu, Y., Weninger, G., Tchagou, C., Marks, A.R.(2024) Nat Commun 15: 8080-8080
- PubMed: 39278969 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-51791-y
- Primary Citation Related Structures:
8UQ2, 8UQ3, 8UQ4, 8UQ5, 8UXC, 8UXE, 8UXF, 8UXG, 8UXH, 8UXI, 8UXL, 8UXM - PubMed Abstract:
Heart failure, the leading cause of mortality and morbidity in the developed world, is characterized by cardiac ryanodine receptor 2 channels that are hyperphosphorylated, oxidized, and depleted of the stabilizing subunit calstabin-2. This results in a diastolic sarcoplasmic reticulum Ca 2+ leak that impairs cardiac contractility and triggers arrhythmias. Genetic mutations in ryanodine receptor 2 can also cause Ca 2+ leak, leading to arrhythmias and sudden cardiac death. Here, we solved the cryogenic electron microscopy structures of ryanodine receptor 2 variants linked either to heart failure or inherited sudden cardiac death. All are in the primed state, part way between closed and open. Binding of Rycal drugs to ryanodine receptor 2 channels reverts the primed state back towards the closed state, decreasing Ca 2+ leak, improving cardiac function, and preventing arrhythmias. We propose a structural-physiological mechanism whereby the ryanodine receptor 2 channel primed state underlies the arrhythmias in heart failure and arrhythmogenic disorders.
- Department of Physiology and Cellular Biophysics, Columbia University Vagelos College of Physicians and Surgeons, New York, NY, USA. mm5642@cumc.columbia.edu.
Organizational Affiliation: