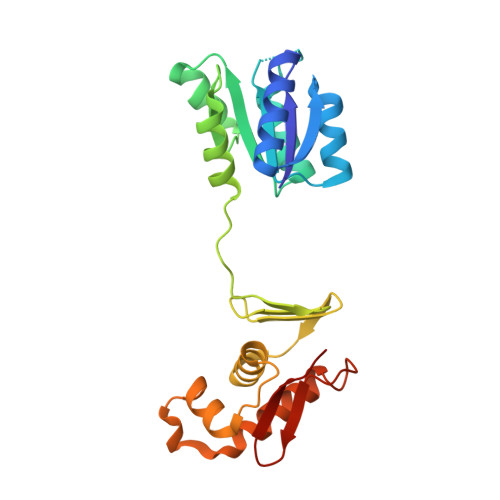
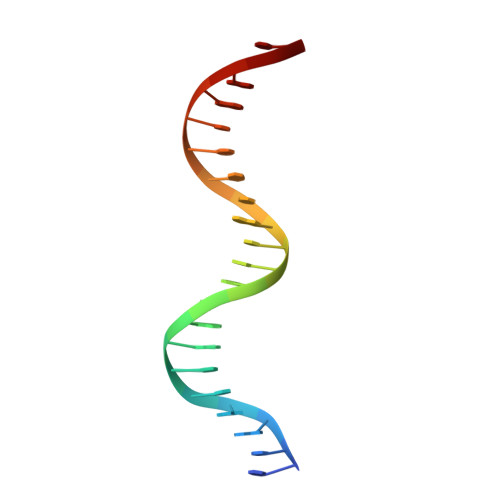
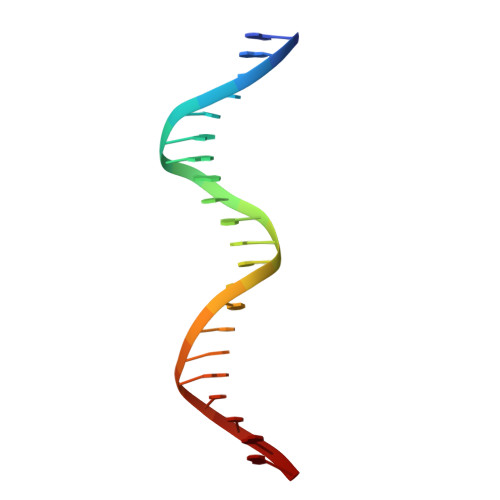
Structural basis of DNA recognition of the Campylobacter jejuni CosR regulator.
Zhang, Z., Yan, Y., Pang, J., Dai, L., Zhang, Q., Yu, E.W.(2024) mBio 15: e0343023-e0343023
- PubMed: 38323832 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mbio.03430-23
- Primary Citation Related Structures:
8UUZ, 8UVK, 8UVX - PubMed Abstract:
Campylobacter jejuni is a foodborne pathogen commonly found in the intestinal tracts of animals. This pathogen is a leading cause of gastroenteritis in humans. Besides its highly infectious nature, C. jejuni is increasingly resistant to a number of clinically administrated antibiotics. As a consequence, the Centers for Disease Control and Prevention has designated antibiotic-resistant Campylobacter as a serious antibiotic resistance threat in the United States. The C. jejuni CosR regulator is essential to the viability of this bacterium and is responsible for regulating the expression of a number of oxidative stress defense enzymes. Importantly, it also modulates the expression of the CmeABC multidrug efflux system, the most predominant and clinically important system in C. jejuni that mediates resistance to multiple antimicrobials. Here, we report structures of apo-CosR and CosR bound with a 21 bp DNA sequence located at the cmeABC promotor region using both single-particle cryo-electron microscopy and X-ray crystallography. These structures allow us to propose a novel mechanism for CosR regulation that involves a long-distance conformational coupling and rearrangement of the secondary structural elements of the regulator to bind target DNA. Campylobacter jejuni has emerged as an antibiotic-resistant threat worldwide. CosR is an essential regulator for this bacterium and is important for Campylobacter adaptation to various stresses. Here, we describe the structural basis of CosR binding to target DNA as determined by cryo-electron microscopy and X-ray crystallography. Since CosR is a potential target for intervention, our studies may facilitate the development of novel therapeutics to combat C. jejuni infection.
- Department of Pharmacology, Case Western Reserve University School of Medicine, Cleveland, Ohio, USA.
Organizational Affiliation: