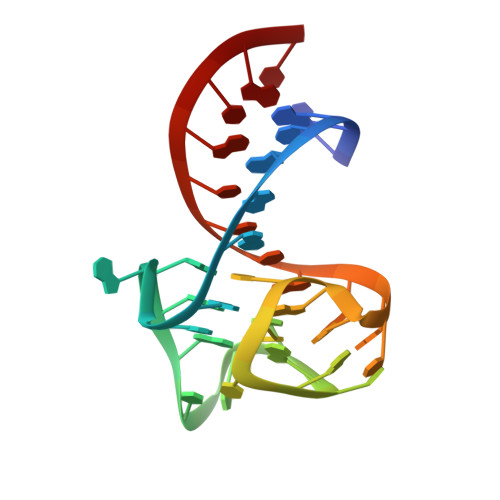
Symmetry breaking of fluorophore binding to a G-quadruplex generates an RNA aptamer with picomolar KD.
Lu, X., Passalacqua, L.F.M., Nodwell, M., Kong, K.Y.S., Caballero-Garcia, G., Dolgosheina, E.V., Ferre-D'Amare, A.R., Britton, R., Unrau, P.J.(2024) Nucleic Acids Res 52: 8039-8051
- PubMed: 38945550 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkae493
- Primary Citation Related Structures:
8U5J, 8U5K, 8U5P, 8U5R, 8U5T, 8U5Z, 8U60 - PubMed Abstract:
Fluorogenic RNA aptamer tags with high affinity enable RNA purification and imaging. The G-quadruplex (G4) based Mango (M) series of aptamers were selected to bind a thiazole orange based (TO1-Biotin) ligand. Using a chemical biology and reselection approach, we have produced a MII.2 aptamer-ligand complex with a remarkable set of properties: Its unprecedented KD of 45 pM, formaldehyde resistance (8% v/v), temperature stability and ligand photo-recycling properties are all unusual to find simultaneously within a small RNA tag. Crystal structures demonstrate how MII.2, which differs from MII by a single A23U mutation, and modification of the TO1-Biotin ligand to TO1-6A-Biotin achieves these results. MII binds TO1-Biotin heterogeneously via a G4 surface that is surrounded by a stadium of five adenosines. Breaking this pseudo-rotational symmetry results in a highly cooperative and homogeneous ligand binding pocket: A22 of the G4 stadium stacks on the G4 binding surface while the TO1-6A-Biotin ligand completely fills the remaining three quadrants of the G4 ligand binding face. Similar optimization attempts with MIII.1, which already binds TO1-Biotin in a homogeneous manner, did not produce such marked improvements. We use the novel features of the MII.2 complex to demonstrate a powerful optically-based RNA purification system.
- Department of Molecular Biology and Biochemistry, Simon Fraser University, Burnaby, British Columbia V5A 1S6, Canada.
Organizational Affiliation: