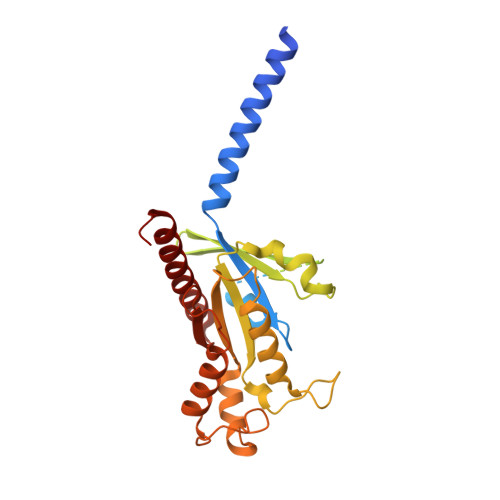
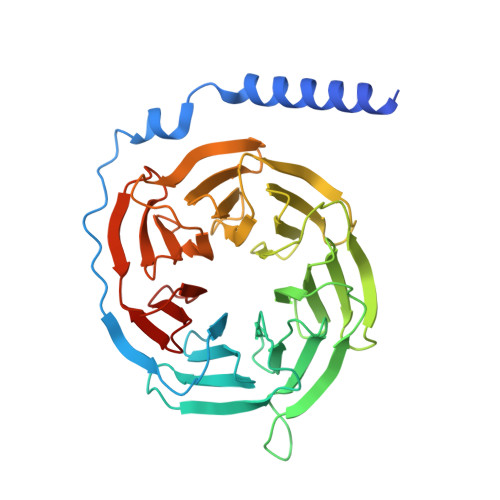
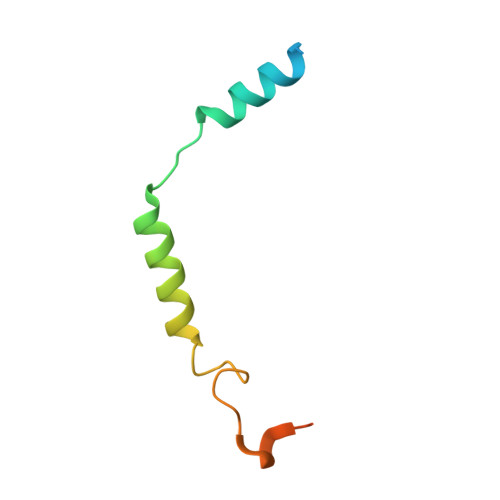
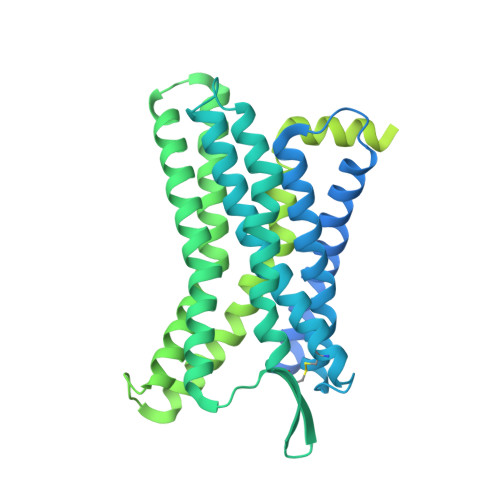
Structural insights into CXCR4 modulation and oligomerization.
Saotome, K., McGoldrick, L.L., Ho, J.H., Ramlall, T.F., Shah, S., Moore, M.J., Kim, J.H., Leidich, R., Olson, W.C., Franklin, M.C.(2025) Nat Struct Mol Biol 32: 315-325
- PubMed: 39313635 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-024-01397-1
- Primary Citation Related Structures:
8U4N, 8U4O, 8U4P, 8U4Q, 8U4R, 8U4S, 8U4T - PubMed Abstract:
Activation of the chemokine receptor CXCR4 by its chemokine ligand CXCL12 regulates diverse cellular processes. Previously reported crystal structures of CXCR4 revealed the architecture of an inactive, homodimeric receptor. However, many structural aspects of CXCR4 remain poorly understood. Here, we use cryo-electron microscopy to investigate various modes of human CXCR4 regulation. CXCL12 activates CXCR4 by inserting its N terminus deep into the CXCR4 orthosteric pocket. The binding of US Food and Drug Administration-approved antagonist AMD3100 is stabilized by electrostatic interactions with acidic residues in the seven-transmembrane-helix bundle. A potent antibody blocker, REGN7663, binds across the extracellular face of CXCR4 and inserts its complementarity-determining region H3 loop into the orthosteric pocket. Trimeric and tetrameric structures of CXCR4 reveal modes of G-protein-coupled receptor oligomerization. We show that CXCR4 adopts distinct subunit conformations in trimeric and tetrameric assemblies, highlighting how oligomerization could allosterically regulate chemokine receptor function.
- Regeneron Pharmaceuticals, Inc., Tarrytown, NY, USA. kei.saotome@regeneron.com.
Organizational Affiliation: