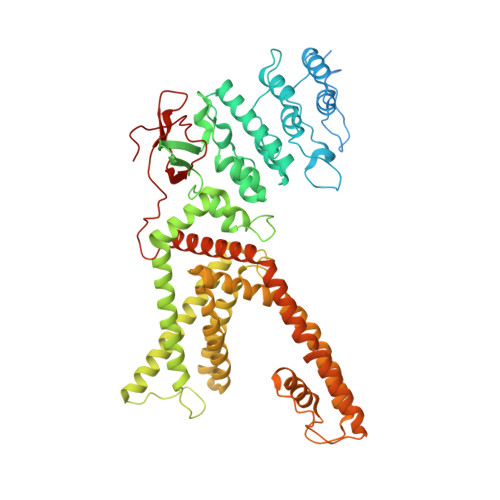
Structural basis of TRPV1 modulation by endogenous bioactive lipids.
Arnold, W.R., Mancino, A., Moss 3rd, F.R., Frost, A., Julius, D., Cheng, Y.(2024) Nat Struct Mol Biol 31: 1377-1385
- PubMed: 38698206 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-024-01299-2
- Primary Citation Related Structures:
8T0C, 8T0E, 8T0Y, 8T10, 8T3L, 8T3M, 8U2Z, 8U30, 8U3A, 8U3C, 8U3J, 8U3L, 8U43, 8U4D - PubMed Abstract:
TRP ion channels are modulated by phosphoinositide lipids, but the underlying structural mechanisms remain unclear. The capsaicin- and heat-activated receptor, TRPV1, has served as a model for deciphering lipid modulation, which is relevant to understanding how pro-algesic agents enhance channel activity in the setting of inflammatory pain. Identification of a pocket within the TRPV1 transmembrane core has provided initial clues as to how phosphoinositide lipids bind to and regulate the channel. Here we show that this regulatory pocket in rat TRPV1 can accommodate diverse lipid species, including the inflammatory lipid lysophosphatidic acid, whose actions are determined by their specific modes of binding. Furthermore, we show that an empty-pocket channel lacking an endogenous phosphoinositide lipid assumes an agonist-like state, even at low temperature, substantiating the concept that phosphoinositide lipids serve as negative TRPV1 modulators whose ejection from the binding pocket is a critical step toward activation by thermal or chemical stimuli.
- Department of Biochemistry and Biophysics, University of California San Francisco, San Francisco, CA, USA.
Organizational Affiliation: