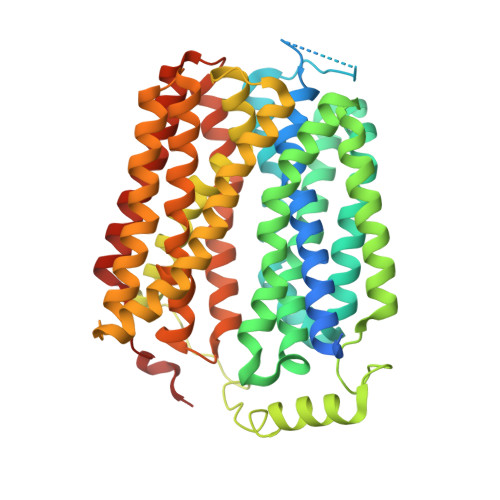
Structure and inhibition of the human lysosomal transporter Sialin.
Schmiege, P., Donnelly, L., Elghobashi-Meinhardt, N., Lee, C.H., Li, X.(2024) Nat Commun 15: 4386-4386
- PubMed: 38782953 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-48535-3
- Primary Citation Related Structures:
8U3D, 8U3E, 8U3F, 8U3G, 8U3H, 9AYB - PubMed Abstract:
Sialin, a member of the solute carrier 17 (SLC17) transporter family, is unique in its ability to transport not only sialic acid using a pH-driven mechanism, but also transport mono and diacidic neurotransmitters, such as glutamate and N-acetylaspartylglutamate (NAAG), into synaptic vesicles via a membrane potential-driven mechanism. While most transporters utilize one of these mechanisms, the structural basis of how Sialin transports substrates using both remains unclear. Here, we present the cryogenic electron-microscopy structures of human Sialin: apo cytosol-open, apo lumen-open, NAAG-bound, and inhibitor-bound. Our structures show that a positively charged cytosol-open vestibule accommodates either NAAG or the Sialin inhibitor Fmoc-Leu-OH, while its luminal cavity potentially binds sialic acid. Moreover, functional analyses along with molecular dynamics simulations identify key residues in binding sialic acid and NAAG. Thus, our findings uncover the essential conformational states in NAAG and sialic acid transport, demonstrating a working model of SLC17 transporters.
- Department of Molecular Genetics, University of Texas Southwestern Medical Center, Dallas, TX, USA.
Organizational Affiliation: