Overcoming resolution attenuation during tilted cryo-EM data collection.
Aiyer, S., Baldwin, P.R., Tan, S.M., Shan, Z., Oh, J., Mehrani, A., Bowman, M.E., Louie, G., Passos, D.O., Dordevic-Marquardt, S., Mietzsch, M., Hull, J.A., Hoshika, S., Barad, B.A., Grotjahn, D.A., McKenna, R., Agbandje-McKenna, M., Benner, S.A., Noel, J.A.P., Wang, D., Tan, Y.Z., Lyumkis, D.(2024) Nat Commun 15: 389-389
- PubMed: 38195598 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-44555-7
- Primary Citation Related Structures:
8TXO - PubMed Abstract:
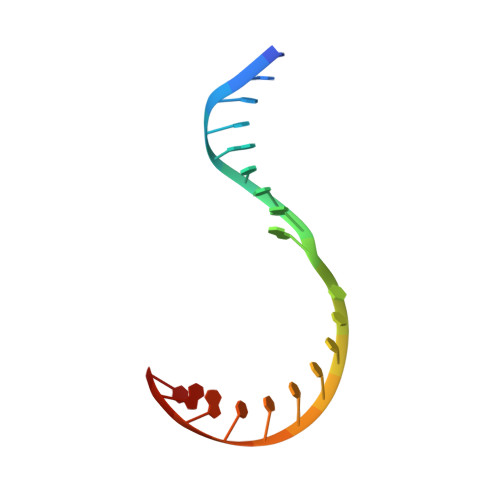
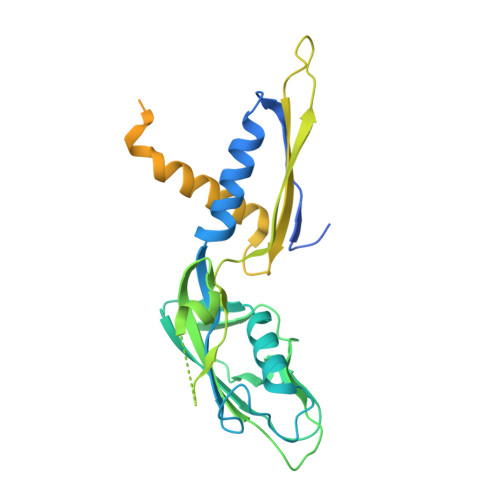
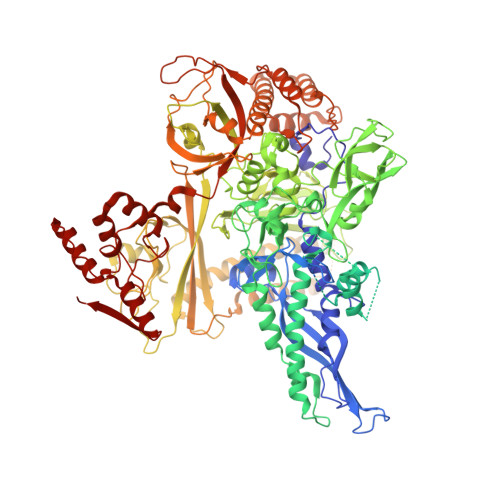
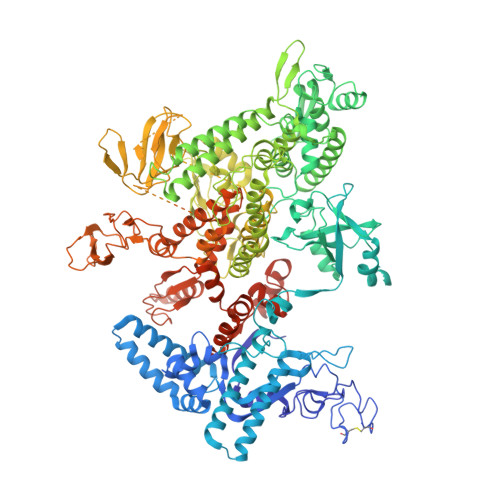
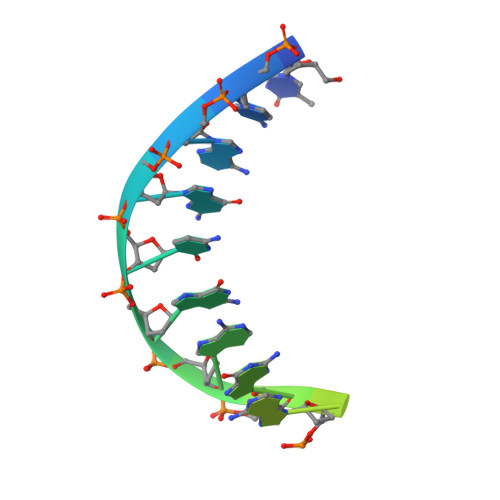
Structural biology efforts using cryogenic electron microscopy are frequently stifled by specimens adopting "preferred orientations" on grids, leading to anisotropic map resolution and impeding structure determination. Tilting the specimen stage during data collection is a generalizable solution but has historically led to substantial resolution attenuation. Here, we develop updated data collection and image processing workflows and demonstrate, using multiple specimens, that resolution attenuation is negligible or significantly reduced across tilt angles. Reconstructions with and without the stage tilted as high as 60° are virtually indistinguishable. These strategies allowed the reconstruction to 3 Å resolution of a bacterial RNA polymerase with preferred orientation, containing an unnatural nucleotide for studying novel base pair recognition. Furthermore, we present a quantitative framework that allows cryo-EM practitioners to define an optimal tilt angle during data acquisition. These results reinforce the utility of employing stage tilt for data collection and provide quantitative metrics to obtain isotropic maps.
- Laboratory of Genetics, The Salk Institute for Biological Studies, La Jolla, CA, 92037, USA.
Organizational Affiliation: