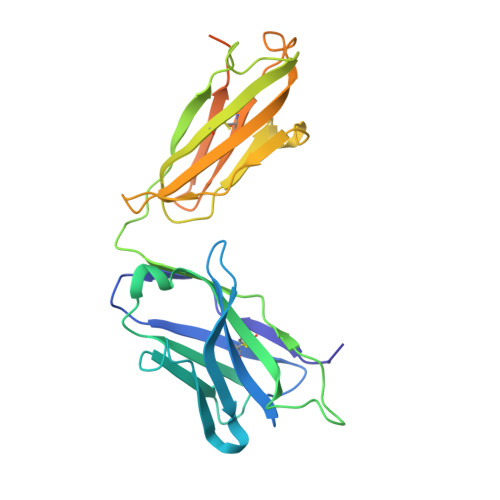
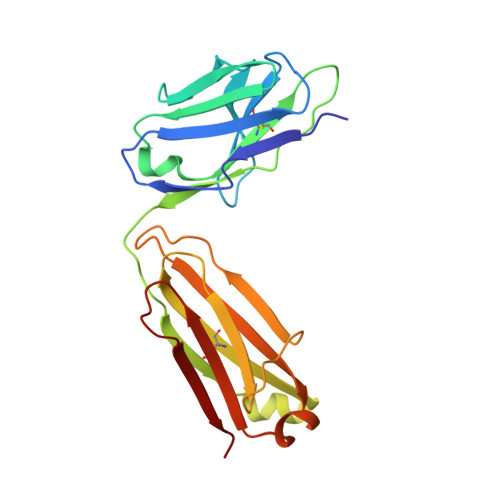
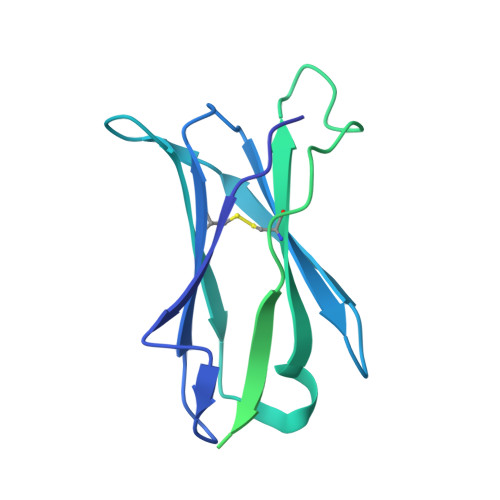
Structural basis for the activity and specificity of the immune checkpoint inhibitor lirilumab.
Lorig-Roach, N., Harpell, N.M., DuBois, R.M.(2024) Sci Rep 14: 742-742
- PubMed: 38185735 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-023-50262-6
- Primary Citation Related Structures:
8TUI - PubMed Abstract:
The clinical success of immune checkpoint inhibitors has underscored the key role of the immune system in controlling cancer. Current FDA-approved immune checkpoint inhibitors target the regulatory receptor pathways of cytotoxic T-cells to enhance their anticancer responses. Despite an abundance of evidence that natural killer (NK) cells can also mediate potent anticancer activities, there are no FDA-approved inhibitors targeting NK cell specific checkpoint pathways. Lirilumab, the most clinically advanced NK cell checkpoint inhibitor, targets inhibitory killer immunoglobulin-like receptors (KIRs), however it has yet to conclusively demonstrate clinical efficacy. Here we describe the crystal structure of lirilumab in complex with the inhibitory KIR2DL3, revealing the precise epitope of lirilumab and the molecular mechanisms underlying KIR checkpoint blockade. Notably, the epitope includes several key amino acids that vary across the human population, and binding studies demonstrate the importance of these amino acids for lirilumab binding. These studies reveal how KIR variations in patients could influence the clinical efficacy of lirilumab and reveal general concepts for the development of immune checkpoint inhibitors targeting NK cells.
- Department of Biomolecular Engineering, University of California Santa Cruz, Santa Cruz, CA, USA.
Organizational Affiliation: