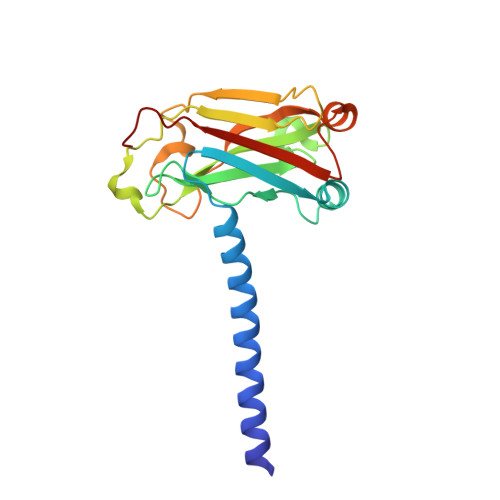
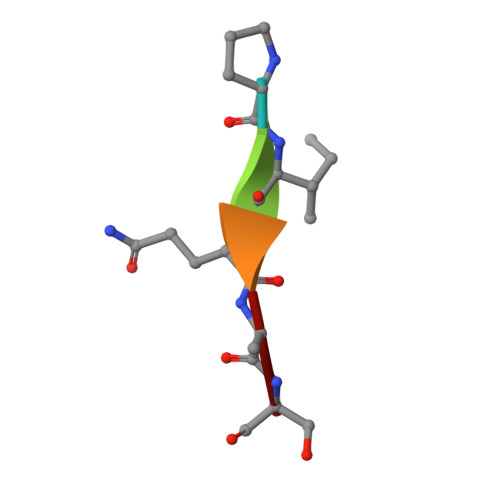
SARS-CoV-2 ORF3a-Mediated NF-kappa B Activation Is Not Dependent on TRAF-Binding Sequence.
Busscher, B.M., Befekadu, H.B., Liu, Z., Xiao, T.S.(2023) Viruses 15
- PubMed: 38005906
- DOI: https://doi.org/10.3390/v15112229
- Primary Citation Related Structures:
8T5P, 8T5Q - PubMed Abstract:
Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) has caused a global pandemic of Coronavirus Disease 2019 (COVID-19). Excessive inflammation is a hallmark of severe COVID-19, and several proteins encoded in the SARS-CoV-2 genome are capable of stimulating inflammatory pathways. Among these, the accessory protein open reading frame 3a (ORF3a) has been implicated in COVID-19 pathology. Here we investigated the roles of ORF3a in binding to TNF receptor-associated factor (TRAF) proteins and inducing nuclear factor kappa B (NF-κB) activation. X-ray crystallography and a fluorescence polarization assay revealed low-affinity binding between an ORF3a N-terminal peptide and TRAFs, and a dual-luciferase assay demonstrated NF-κB activation by ORF3a. Nonetheless, mutation of the N-terminal TRAF-binding sequence PIQAS in ORF3a did not significantly diminish NF-κB activation in our assay. Our results thus suggest that the SARS-CoV-2 protein may activate NF-κB through alternative mechanisms.
- Department of Pathology, School of Medicine, Case Western Reserve University, Cleveland, OH 44106, USA.
Organizational Affiliation: