Cryo-EM structure of the band 3 anion transport protein
Su, C.-C.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Band 3 anion transport protein | 911 | Homo sapiens | Mutation(s): 0 Membrane Entity: Yes | 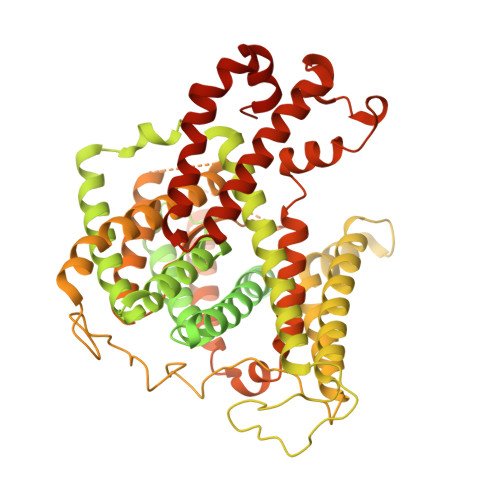 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P02730 GTEx: ENSG00000004939 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P02730 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: P02730-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PIO Download:Ideal Coordinates CCD File | E [auth A], F [auth B] | [(2R)-2-octanoyloxy-3-[oxidanyl-[(1R,2R,3S,4R,5R,6S)-2,3,6-tris(oxidanyl)-4,5-diphosphonooxy-cyclohexyl]oxy-phosphoryl]oxy-propyl] octanoate C25 H49 O19 P3 XLNCEHRXXWQMPK-MJUMVPIBSA-N |  | ||
| PLC Download:Ideal Coordinates CCD File | D [auth A], G [auth B] | DIUNDECYL PHOSPHATIDYL CHOLINE C32 H65 N O8 P IJFVSSZAOYLHEE-SSEXGKCCSA-O |  | ||
| NAG Download:Ideal Coordinates CCD File | C [auth A], H [auth B] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute Of Allergy and Infectious Diseases (NIH/NIAID) | United States | -- |