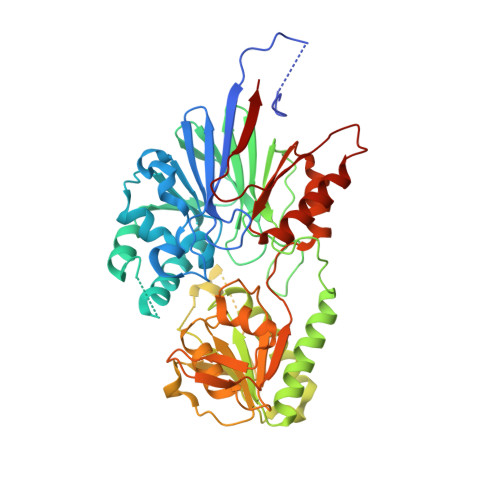
Anticancer benzoxaboroles block pre-mRNA processing by directly inhibiting CPSF3.
Tao, Y., Budhipramono, A., Huang, J., Fang, M., Xie, S., Kim, J., Khivansara, V., Dominski, Z., Tong, L., De Brabander, J.K., Nijhawan, D.(2024) Cell Chem Biol 31: 139
- PubMed: 37967558 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chembiol.2023.10.019
- Primary Citation Related Structures:
8T1Q, 8T1R - PubMed Abstract:
A novel class of benzoxaboroles was reported to induce cancer cell death but the mechanism was unknown. Using a forward genetics platform, we discovered mutations in cleavage and polyadenylation specific factor 3 (CPSF3) that reduce benzoxaborole binding and confer resistance. CPSF3 is the endonuclease responsible for pre-mRNA 3'-end processing, which is also important for RNA polymerase II transcription termination. Benzoxaboroles inhibit this endonuclease activity of CPSF3 in vitro and also curb transcriptional termination in cells, which results in the downregulation of numerous constitutively expressed genes. Furthermore, we used X-ray crystallography to demonstrate that benzoxaboroles bind to the active site of CPSF3 in a manner distinct from the other known inhibitors of CPSF3. The benzoxaborole compound impeded the growth of cancer cell lines derived from different lineages. Our results suggest benzoxaboroles may represent a promising lead as CPSF3 inhibitors for clinical development.
- Department of Biochemistry, University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: