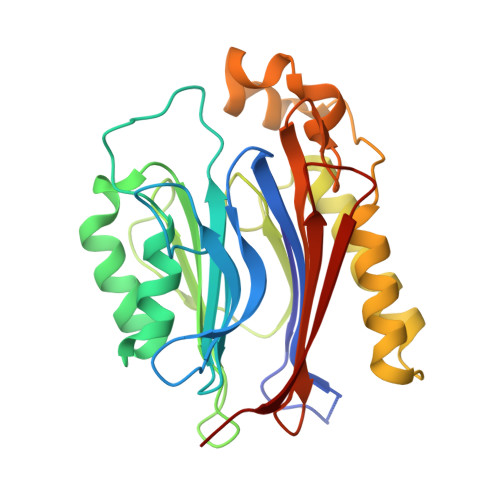
The crystal structure of human transport and Golgi organization 2 homolog (TANGO2) suggests a cysteine N-terminal nucleophile (Ntn) hydrolase.
Zhou, D., Chen, L., Rose, J., Wang, B.C.(2026) Acta Crystallogr D Struct Biol 82: 383-396
- PubMed: 41924852 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798326001968
- Primary Citation Related Structures:
8SV7 - PubMed Abstract:
Recently, there has been growing interest in the function and physiological importance of human TANGO2 (transport and Golgi organization 2 homolog), particularly whether it acts as a heme-trafficking protein. To address this question, we experimentally determined the three-dimensional structure of TANGO2. Our crystallographic analysis indicates that interactions between heme and TANGO2 are nonspecific. Structural comparison of the TANGO2 crystal structure with known cysteine Ntn-hydrolases allowed us to identify a putative active site, catalytic residues and a substrate-binding cavity that correspond to residues that are mutated in pathogenic TANGO2 variants. Based on these features, we propose that TANGO2 may utilize fatty-acid derivatives as substrates, suggesting a potential role in lipid metabolism. Mutations in the human TANGO2 gene cause TANGO2 deficiency disorder, a multisystem, life-threatening disease with onset in early childhood. Together, our results provide new insights into the molecular function of TANGO2 and help to resolve the ongoing debate regarding whether it functions as a heme-trafficking protein.
- Department of Biochemistry and Molecular Biology, University of Georgia, Athens, GA 30602, USA.
Organizational Affiliation: