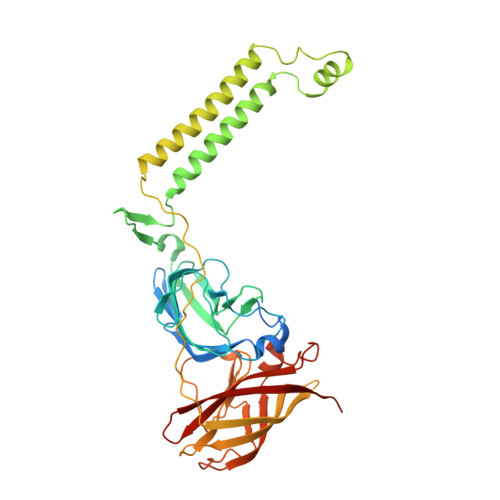
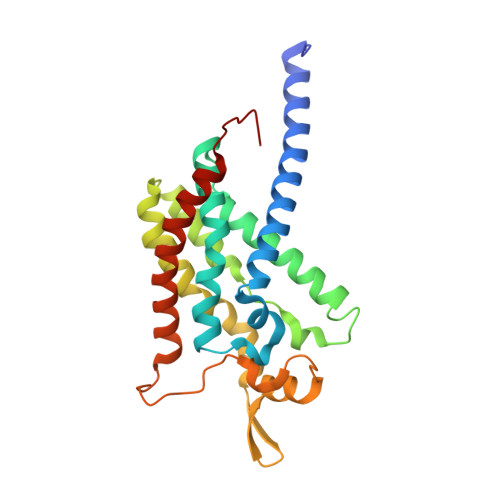
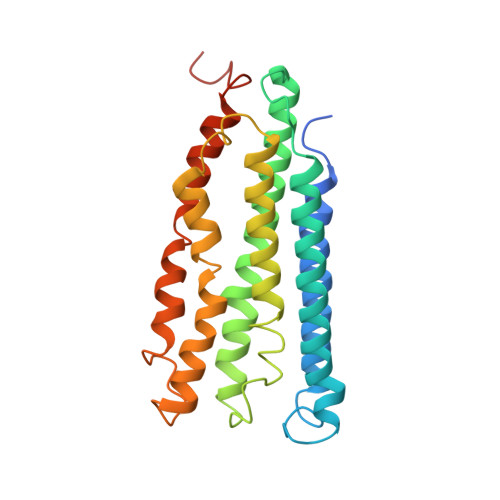
Product analog binding identifies the copper active site of particulate methane monooxygenase.
Tucci, F.J., Jodts, R.J., Hoffman, B.M., Rosenzweig, A.C.(2023) Nat Catal 6: 1194-1204
- PubMed: 38187819 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41929-023-01051-x
- Primary Citation Related Structures:
8OYI, 8SQW, 8SR1, 8SR2, 8SR4, 8SR5 - PubMed Abstract:
Nature's primary methane-oxidizing enzyme, the membrane-bound particulate methane monooxygenase (pMMO), catalyzes the oxidation of methane to methanol. pMMO activity requires copper, and decades of structural and spectroscopic studies have sought to identify the active site among three candidates: the Cu B , Cu C , and Cu D sites. Challenges associated with the isolation of active pMMO have hindered progress toward locating its catalytic center. However, reconstituting pMMO into native lipid nanodiscs stabilizes its structure and recovers its activity. Here, these active samples were incubated with 2,2,2,-trifluoroethanol (TFE), a product analog that serves as a readily visualized active-site probe. Interactions of TFE with the Cu D site were observed by both pulsed ENDOR spectroscopy and cryoEM, implicating Cu D and the surrounding hydrophobic pocket as the likely site of methane oxidation. Use of these orthogonal techniques on parallel samples is a powerful approach that can circumvent difficulties in interpreting metalloenzyme cryoEM maps.
- Departments of Molecular Biosciences and of Chemistry, Northwestern University, Evanston, IL, USA.
Organizational Affiliation: