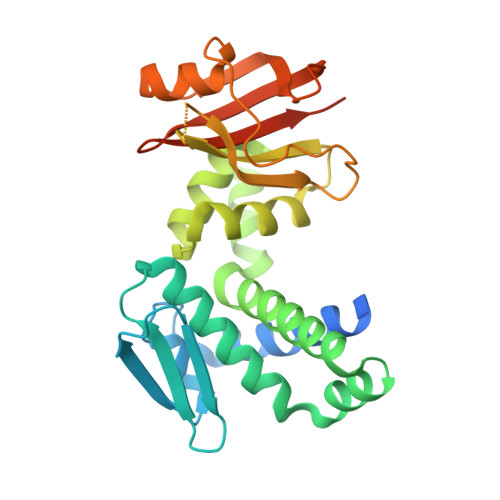
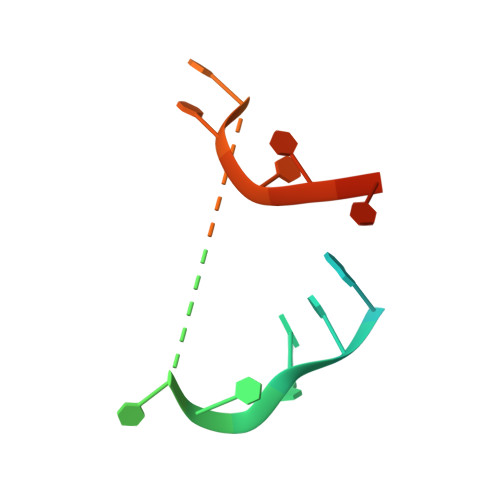
The Deinococcus protease PprI senses DNA damage by directly interacting with single-stranded DNA.
Lu, H., Chen, Z., Xie, T., Zhong, S., Suo, S., Song, S., Wang, L., Xu, H., Tian, B., Zhao, Y., Zhou, R., Hua, Y.(2024) Nat Commun 15: 1892-1892
- PubMed: 38424107 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-46208-9
- Primary Citation Related Structures:
8SLM, 8SLN - PubMed Abstract:
Bacteria have evolved various response systems to adapt to environmental stress. A protease-based derepression mechanism in response to DNA damage was characterized in Deinococcus, which is controlled by the specific cleavage of repressor DdrO by metallopeptidase PprI (also called IrrE). Despite the efforts to document the biochemical, physiological, and downstream regulation of PprI-DdrO, the upstream regulatory signal activating this system remains unclear. Here, we show that single-stranded DNA physically interacts with PprI protease, which enhances the PprI-DdrO interactions as well as the DdrO cleavage in a length-dependent manner both in vivo and in vitro. Structures of PprI, in its apo and complexed forms with single-stranded DNA, reveal two DNA-binding interfaces shaping the cleavage site. Moreover, we show that the dynamic monomer-dimer equilibrium of PprI is also important for its cleavage activity. Our data provide evidence that single-stranded DNA could serve as the signal for DNA damage sensing in the metalloprotease/repressor system in bacteria. These results also shed light on the survival and acquired drug resistance of certain bacteria under antimicrobial stress through a SOS-independent pathway.
- MOE Key Laboratory of Biosystems Homeostasis & Protection, Institute of Biophysics, College of Life Sciences, Zhejiang University, Hangzhou, China.
Organizational Affiliation: