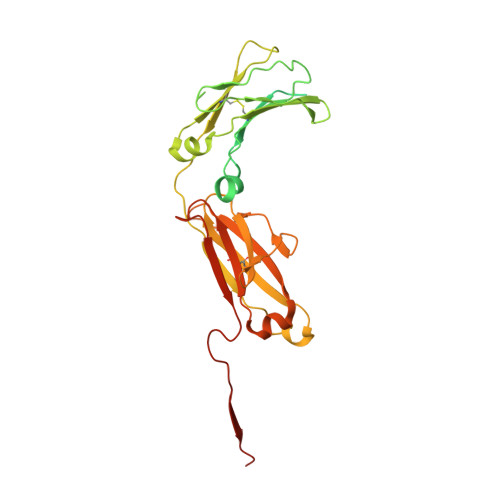
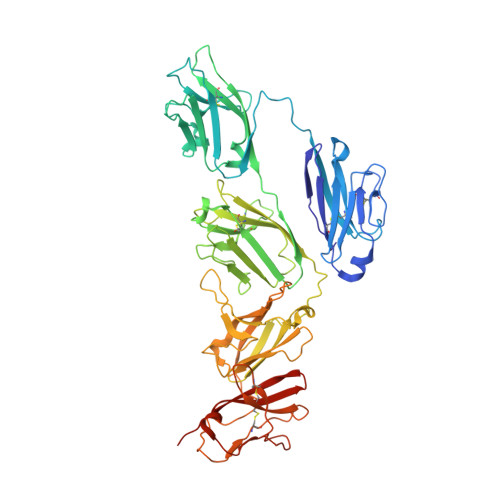
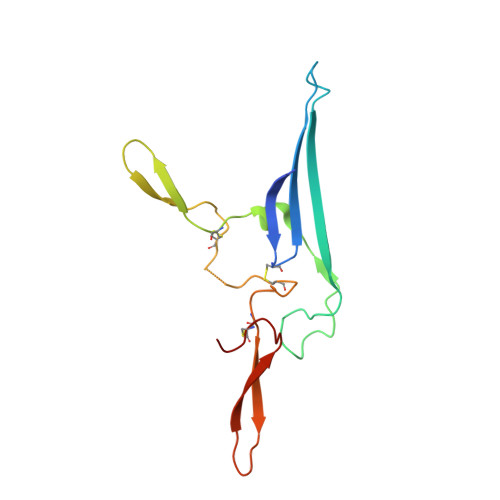
SIgA structures bound to Streptococcus pyogenes M4 and human CD89 provide insights into host-pathogen interactions.
Liu, Q., Stadtmueller, B.M.(2023) Nat Commun 14: 6726-6726
- PubMed: 37872175 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-42469-y
- Primary Citation Related Structures:
8SKU, 8SKV - PubMed Abstract:
Immunoglobulin (Ig) A functions as monomeric IgA in the serum and Secretory (S) IgA in mucosal secretions. Host IgA Fc receptors (FcαRs), including human FcαR1/CD89, mediate IgA effector functions; however, human pathogen Streptococcus pyogenes has evolved surface-protein virulence factors, including M4, that also engage the CD89-binding site on IgA. Despite human mucosa serving as a reservoir for pathogens, SIgA interactions with CD89 and M4 remain poorly understood. Here we report cryo-EM structures of M4-SIgA and CD89-SIgA complexes, which unexpectedly reveal different SIgA-binding stoichiometry for M4 and CD89. Structural data, supporting experiments, and modeling indicate that copies of SIgA bound to S. pyogenes M4 will adopt similar orientations on the bacterium surface and leave one host FcαR binding site open. Results suggest unappreciated functional consequences associated with SIgA binding to host and bacterial FcαRs relevant to understanding host-microbe co-evolution, IgA effector functions and improving the outcomes of group A Streptococcus infection.
- Department of Biochemistry, University of Illinois Urbana-Champaign, Urbana, Illinois, 61801, USA.
Organizational Affiliation: