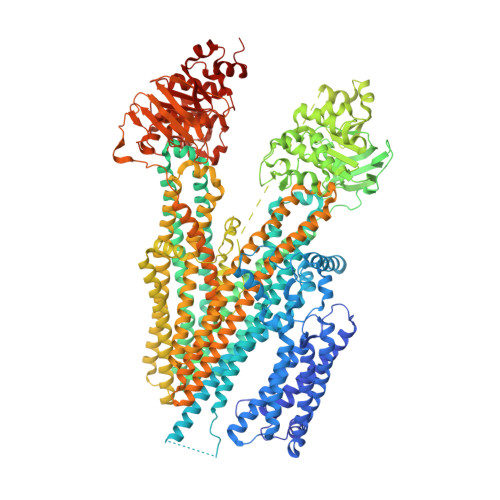
Structural basis for autoinhibition by the dephosphorylated regulatory domain of Ycf1.
Khandelwal, N.K., Tomasiak, T.M.(2024) Nat Commun 15: 2389-2389
- PubMed: 38493146 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-46722-w
- Primary Citation Related Structures:
8SG4 - PubMed Abstract:
Yeast Cadmium Factor 1 (Ycf1) sequesters glutathione and glutathione-heavy metal conjugates into yeast vacuoles as a cellular detoxification mechanism. Ycf1 belongs to the C subfamily of ATP Binding Cassette (ABC) transporters characterized by long flexible linkers, notably the regulatory domain (R-domain). R-domain phosphorylation is necessary for activity, whereas dephosphorylation induces autoinhibition through an undefined mechanism. Because of its transient and dynamic nature, no structure of the dephosphorylated Ycf1 exists, limiting understanding of this R-domain regulation. Here, we capture the dephosphorylated Ycf1 using cryo-EM and show that the unphosphorylated R-domain indeed forms an ordered structure with an unexpected hairpin topology bound within the Ycf1 substrate cavity. This architecture and binding mode resemble that of a viral peptide inhibitor of an ABC transporter and the secreted bacterial WXG peptide toxins. We further reveal the subset of phosphorylation sites within the hairpin turn that drive the reorganization of the R-domain conformation, suggesting a mechanism for Ycf1 activation by phosphorylation-dependent release of R-domain mediated autoinhibition.
- Department of Chemistry and Biochemistry, University of Arizona, Tucson, AZ, 85721, USA.
Organizational Affiliation: