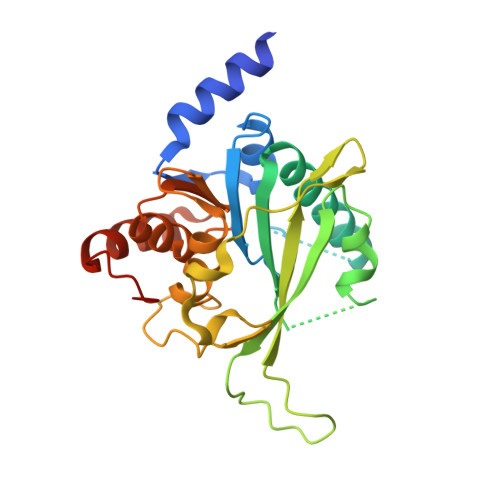
Biochemical and structural characterization of the first-discovered metazoan DNA cytosine-N4 methyltransferase from the bdelloid rotifer Adineta vaga.
Zhou, J., Horton, J.R., Kaur, G., Chen, Q., Li, X., Mendoza, F., Wu, T., Blumenthal, R.M., Zhang, X., Cheng, X.(2023) J Biological Chem 299: 105017-105017
- PubMed: 37414145
- DOI: https://doi.org/10.1016/j.jbc.2023.105017
- Primary Citation of Related Structures:
8S9M, 8S9N, 8S9O - PubMed Abstract:
Much is known about the generation, removal, and roles of 5-methylcytosine (5mC) in eukaryote DNA, and there is a growing body of evidence regarding N6-methyladenine, but very little is known about N4-methylcytosine (4mC) in the DNA of eukaryotes. The gene for the first metazoan DNA methyltransferase generating 4mC (N4CMT) was reported and characterized recently by others, in tiny freshwater invertebrates called bdelloid rotifers. Bdelloid rotifers are ancient, apparently asexual animals, and lack canonical 5mC DNA methyltransferases. Here, we characterize the kinetic properties and structural features of the catalytic domain of the N4CMT protein from the bdelloid rotifer Adineta vaga. We find that N4CMT generates high-level methylation at preferred sites, (a/c)CG(t/c/a), and low-level methylation at disfavored sites, exemplified by ACGG. Like the mammalian de novo 5mC DNA methyltransferase 3A/3B (DNMT3A/3B), N4CMT methylates CpG dinucleotides on both DNA strands, generating hemimethylated intermediates and eventually fully methylated CpG sites, particularly in the context of favored symmetric sites. In addition, like DNMT3A/3B, N4CMT methylates non-CpG sites, mainly CpA/TpG, though at a lower rate. Both N4CMT and DNMT3A/3B even prefer similar CpG-flanking sequences. Structurally, the catalytic domain of N4CMT closely resembles the Caulobacter crescentus cell cycle-regulated DNA methyltransferase. The symmetric methylation of CpG, and similarity to a cell cycle-regulated DNA methyltransferase, together suggest that N4CMT might also carry out DNA synthesis-dependent methylation following DNA replication.
- Department of Epigenetics and Molecular Carcinogenesis, University of Texas MD Anderson Cancer Center, Houston, Texas, USA.
Organizational Affiliation: