Structure of UP1 S4ES6E phosphomimetic mutant in complex with human telomeric repeat DNA
Dunnett, L., Prischi, F.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Heterogeneous nuclear ribonucleoprotein A1, N-terminally processed | 198 | Homo sapiens | Mutation(s): 2 Gene Names: HNRNPA1, HNRPA1 | 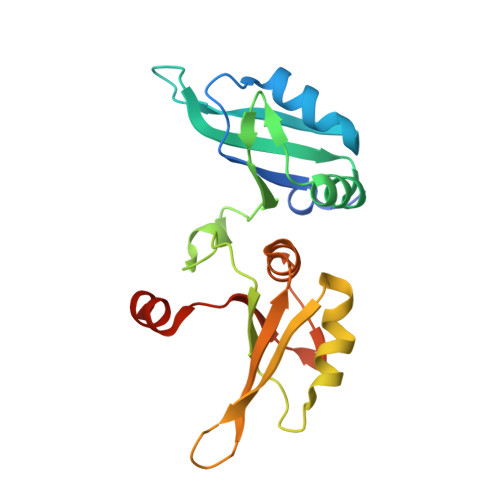 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P09651 (Homo sapiens) Explore P09651 Go to UniProtKB: P09651 | |||||
PHAROS: P09651 GTEx: ENSG00000135486 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P09651 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar nucleic acids by: Sequence
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | Organism | Image | |
| DNA (5'-D(P*TP*AP*GP*GP*GP*TP*TP*AP*GP*GP*G)-3') | 12 | Homo sapiens | 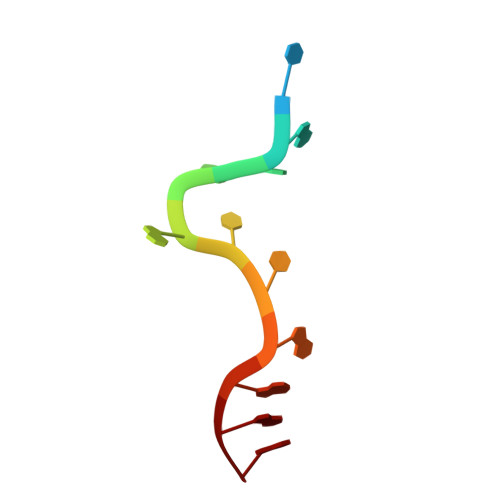 | ||
Sequence AnnotationsExpand | |||||
| |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 51.12 | α = 90 |
| b = 51.12 | β = 90 |
| c = 173.27 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| xia2 | data reduction |
| Aimless | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |