Characterisation of novel influenza-derived HLA-B*18:01-restricted epitopes.
Leong, S.L., Murdolo, L., Maddumage, J.C., Koutsakos, M., Kedzierska, K., Purcell, A.W., Gras, S., Grant, E.J.(2024) Clin Transl Immunology 13: e1509-e1509
- PubMed: 38737448 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cti2.1509
- Primary Citation Related Structures:
8RNG, 8RNH, 8ROO, 8ROP - PubMed Abstract:
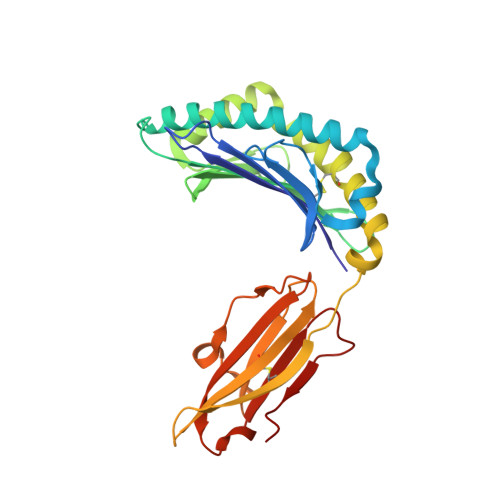
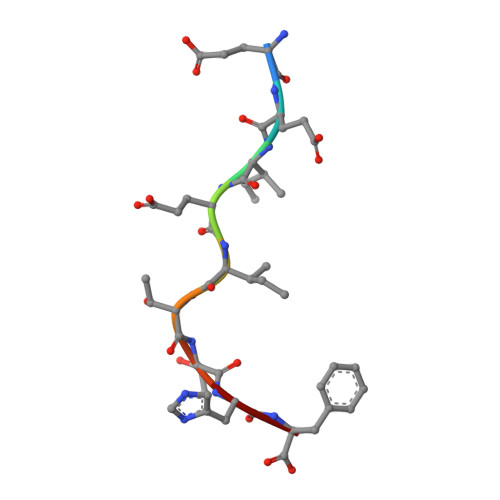
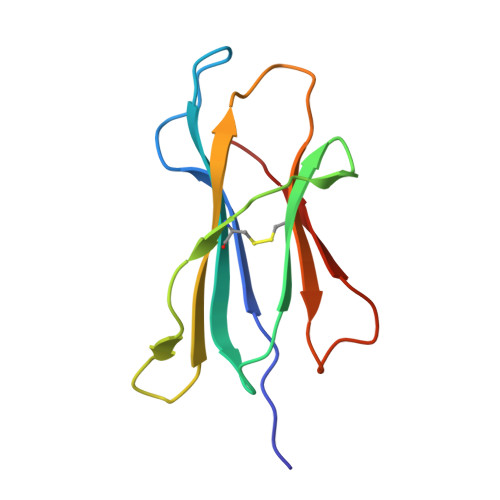
Seasonal influenza viruses cause roughly 650 000 deaths annually despite available vaccines. CD8 + T cells typically recognise influenza-derived peptides from internal structural and non-structural influenza proteins and are an attractive avenue for future vaccine design as they could reduce the severity of disease following infection with diverse influenza strains. CD8 + T cells recognise peptides presented by the highly polymorphic Human Leukocyte Antigens class I molecules (HLA-I). Each HLA-I variant has distinct peptide binding preferences, representing a significant obstacle for designing vaccines that elicit CD8 + T cell responses across broad populations. Consequently, the rational design of a CD8 + T cell-mediated vaccine would require the identification of highly immunogenic peptides restricted to a range of different HLA molecules. Here, we assessed the immunogenicity of six recently published novel influenza-derived peptides identified by mass-spectrometry and predicted to bind to the prevalent HLA-B*18:01 molecule. Using CD8 + T cell activation assays and protein biochemistry, we showed that 3/6 of the novel peptides were immunogenic in several HLA-B*18:01 + individuals and confirmed their HLA-B*18:01 restriction. We subsequently compared CD8 + T cell responses towards the previously identified highly immunogenic HLA-B*18:01-restricted NP 219 peptide. Using X-ray crystallography, we solved the first crystal structures of HLA-B*18:01 presenting immunogenic influenza-derived peptides. Finally, we dissected the first TCR repertoires specific for HLA-B*18:01 restricted pathogen-derived peptides, identifying private and restricted repertoires against each of the four peptides. Overall the characterisation of these novel immunogenic peptides provides additional HLA-B*18:01-restricted vaccine targets derived from the Matrix protein 1 and potentially the non-structural protein and the RNA polymerase catalytic subunit of influenza viruses.
- Infection and Immunity Program, La Trobe Institute for Molecular Science (LIMS) La Trobe University Bundoora VIC Australia.
Organizational Affiliation: