Divergent mechanisms of steroid inhibition in the human rho1 GABAA receptor
Chen, F., John, C., Rebecca, J.H., Erik, L.(2024) Nat Commun
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
(2024) Nat Commun
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Gamma-aminobutyric acid receptor subunit rho-1 | A, B [auth E], C [auth B], D [auth C], E [auth D] | 479 | Homo sapiens | Mutation(s): 0 Gene Names: GABRR1 Membrane Entity: Yes | 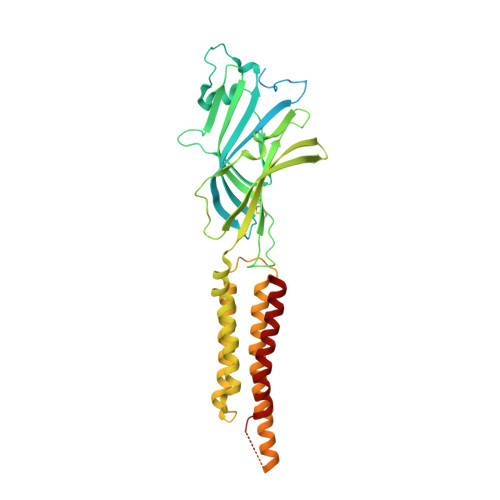 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P24046 GTEx: ENSG00000146276 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P24046 | ||||
Glycosylation | |||||
| Glycosylation Sites: 1 | Go to GlyGen: P24046-1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| EST Download:Ideal Coordinates CCD File | CA [auth E], M [auth A], MA [auth B], NB [auth D], YA [auth C] | ESTRADIOL C18 H24 O2 VOXZDWNPVJITMN-ZBRFXRBCSA-N |  | ||
| NAG (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | F [auth A] FA [auth B] G [auth A] GA [auth B] GB [auth D] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| OCT Download:Ideal Coordinates CCD File | AA [auth E] AB [auth C] BB [auth C] CB [auth D] DB [auth D] | N-OCTANE C8 H18 TVMXDCGIABBOFY-UHFFFAOYSA-N |  | ||
| ABU Download:Ideal Coordinates CCD File | DA [auth E], EA [auth B], FB [auth D], N [auth A], QA [auth C] | GAMMA-AMINO-BUTANOIC ACID C4 H9 N O2 BTCSSZJGUNDROE-UHFFFAOYSA-N |  | ||
| HEX Download:Ideal Coordinates CCD File | BA [auth E] I [auth A] IA [auth B] J [auth A] JA [auth B] | HEXANE C6 H14 VLKZOEOYAKHREP-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | R [auth A] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Swedish Research Council | Sweden | -- |