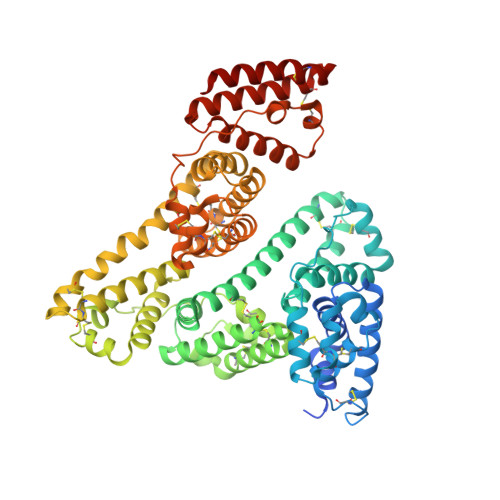
Structural and mechanistic insights into the transport of aristolochic acids and their active metabolites by human serum albumin.
Pomyalov, S., Minetti, C.A., Remeta, D.P., Bonala, R., Johnson, F., Zaitseva, I., Iden, C., Golebiewska, U., Breslauer, K.J., Shoham, G., Sidorenko, V.S., Grollman, A.P.(2024) J Biological Chem 300: 107358-107358
- PubMed: 38782206 Search on PubMed
- DOI: https://doi.org/10.1016/j.jbc.2024.107358
- Primary Citation Related Structures:
8RCO, 8RCP, 8RGK, 8RGL - PubMed Abstract:
Aristolochic acids I and II (AA-I/II) are carcinogenic principles of Aristolochia plants, which have been employed in traditional medicinal practices and discovered as food contaminants. While the deleterious effects of AAs are broadly acknowledged, there is a dearth of information to define the mechanisms underlying their carcinogenicity. Following bioactivation in the liver, N-hydroxyaristolactam and N-sulfonyloxyaristolactam metabolites are transported via circulation and elicit carcinogenic effects by reacting with cellular DNA. In this study, we apply DNA adduct analysis, X-ray crystallography, isothermal titration calorimetry (ITC) and fluorescence quenching to investigate the role of human serum albumin (HSA) in modulating AA carcinogenicity. We find that HSA extends the half-life and reactivity of N-sulfonyloxyaristolactam-I with DNA, thereby protecting activated AAs from heterolysis. Applying novel pooled plasma HSA crystallization methods, we report high-resolution structures of myristic acid-enriched HSA (HSA MYR ) and its AA complexes (HSA MYR /AA-I and HSA MYR /AA-II) at 1.9 Å resolution. Whereas AA-I is located within HSA subdomain IB, AA-II occupies subdomains IIA and IB. ITC binding profiles reveal two distinct AA sites in both complexes with association constants of 1.5 and 0.5 · 10 6 M -1 for HSA/AA-I versus 8.4 and 9.0 · 10 5 M -1 for HSA/AA-II. Fluorescence quenching of the HSA Trp 214 suggests variable impacts of fatty acids on ligand binding affinities. Collectively, our structural and thermodynamic characterizations yield significant insights into AA binding, transport, toxicity, and potential allostery, critical determinants for elucidating the mechanistic roles of HSA in modulating AA carcinogenicity.
- Institute of Chemistry, The Hebrew University of Jerusalem, Edmond J. Safra Campus - Givat Ram, Jerusalem 91904 Israel.
Organizational Affiliation: